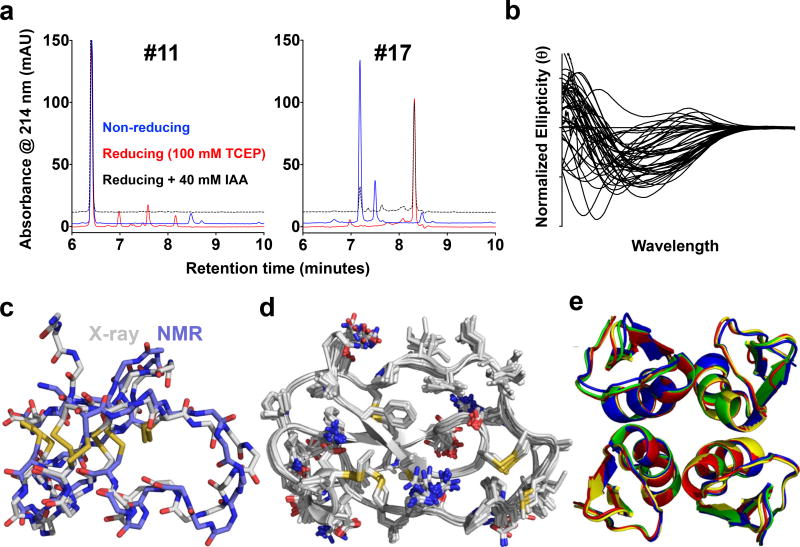
Figure 2. CDP biochemical and crystallographic analyses.
a, Example comparative RPC analyses of two scorpion potassium channel toxins, CDP #11 (left) and #17 (right), produced using the Daedalus-based biologic production platform, under non-reducing conditions (blue), reducing conditions (red; by addition of 100 mM tris(2-carboxyethyl)phosphine (TCEP)), and thiol-blocking treatment after reduction (dashed; by addition of 40 mM iodoacetamide (IAA) after addition of TCEP). b, Normalized CD spectra from all 46 successfully-expressed CDPs showed a wide range of secondary structure compositions. c, A superposition of two structures of CTX (target #28), shown as licorice-stick representations of the peptide backbone plus cysteine side-chains, colored by atom type, determined by NMR (1CHL.pdb33, periwinkle carbons) or by crystallography (reported here, gray carbons). d, A superposition of all 18 independent views of the structure of Elafin (target #4, a human class 1–3, 2–5, 4–6 non-knotted CDP) in the crystal structure reported here, shown as a cartoon ribbon representation of the peptide backbone plus licorice-stick representations of all side-chains, colored by atom type. The structure of Elafin had been previously determined by crystallography (1FLE.pdb34), though only in complex with elastase, and alone by NMR (2REL.pdb35). e, A superposition of all five independent views of the tetramer of the potassium channel toxin gamma-KTx 2.2 hitchin from the venom of the Manchurian scorpion (target #48), from two different crystal forms, shown in a cartoon representation, colored by tetramer.