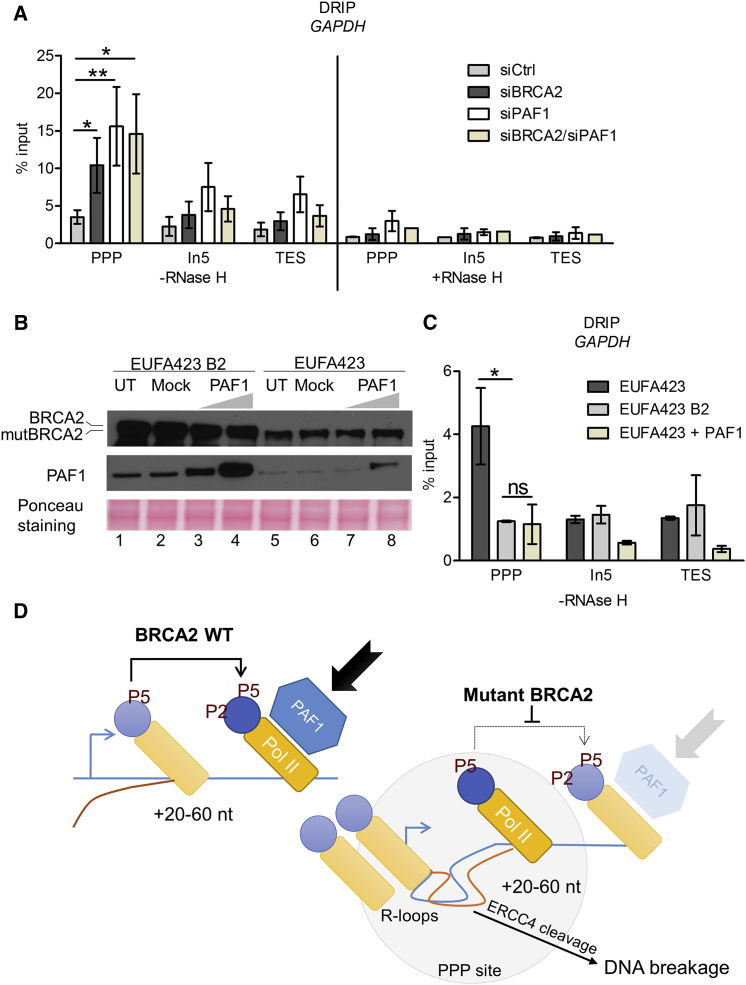
Figure 6.
PAF1 Depletion Phenocopies, while PAF1 Overexpression Ameliorates, R-Loop Accumulation following BRCA2 Inactivation
(A) DRIP analysis with S9.6 antibody of GAPDH gene in HeLa Kyoto cells transfected with indicated siRNAs for 72 hr. R-loop dissolution by RNase H enzyme is shown as a control. Plots show the mean ± SEM from three independent experiments. The 2-way ANOVA test was performed for all pairwise comparisons to determine statistical significance. Statistically significant differences are indicated (∗p < 0.05 and ∗∗p < 0.01).
(B) Western blot analysis of the overexpression of PAF1 in EUFA423 B2 and EUFA423 cells transfected with either 5 μg (lanes 3 and 7) or 10 μg (lanes 4 and 8) of the PAF1 construct. Cell lysates were analyzed 24 hr after transfection. Bars indicate the relative positions of the wild-type and truncated forms of BRCA2 (mutBRCA2).
(C) DRIP analysis of GAPDH gene (as in A) showing R-loop reduction after PAF1 overexpression for 24 hr in EUFA423 cells compared to untransfected EUFA423 or EUFA423 B2 controls. Plots show the mean ± SEM from three independent experiments. The 2-way ANOVA test was performed for all pairwise comparisons to determine statistical significance. Statistically significant differences are indicated (∗p < 0.05; ns, not significant).
(D) Schematic model depicting that BRCA2 regulates transcription elongation by RNA polymerase II to prevent R-loop accumulation. Normally, BRCA2 augments the recruitment of PAF1 to RNAPII paused at PPP sites located ∼20–60 nt from transcription start sites (arrow), fostering the switch to productive transcription elongation (left). This function is subverted by cancer-causing BRCA2 mutations (right), diminishing PAF1 recruitment. Consequently, RNAPII accumulates at PPP sites, triggering the site-specific accrual of unscheduled R-loops, which are cleaved by the ERCC4 endonuclease to generate DNA breaks, triggering genomic instability and carcinogenesis following BRCA2 inactivation.