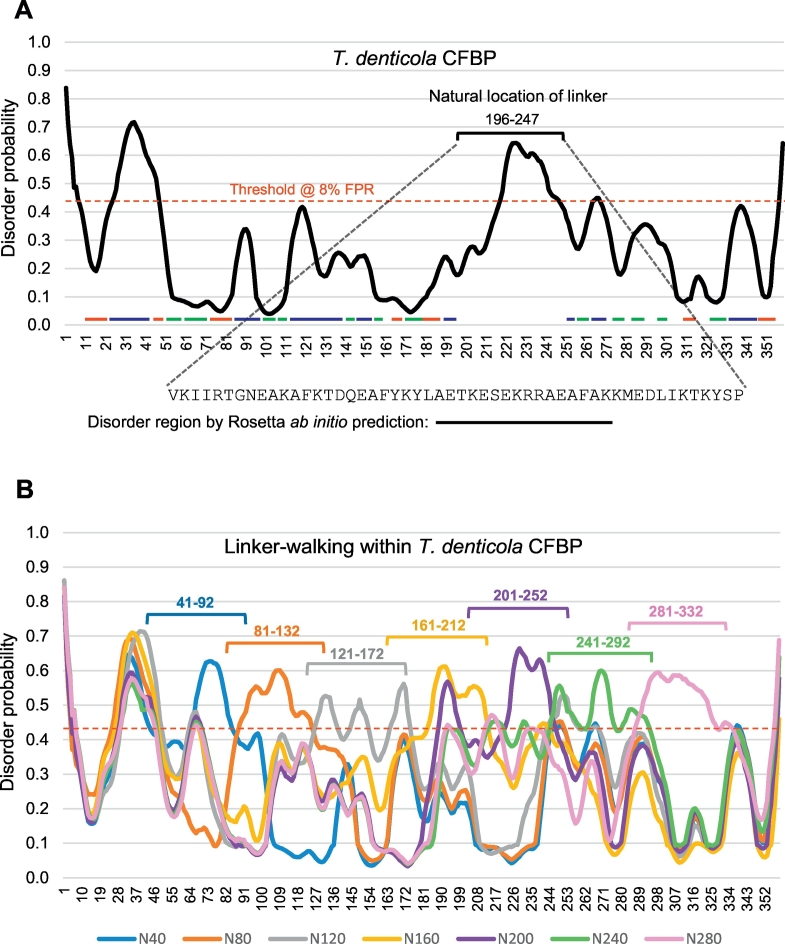
Fig. 6.
Disorder plot for Treponema denticola CFBP with relocated linker sequences. Disorder analysis and plot were performed essentially as in the two preceding Figures (Fig. 4, Fig. 5). (A) Top: Plot for native T. denticola CFBP with the 52-amino acid long linker region in its natural location. In addition, secondary structural elements, corresponding to known structures of homologous CYN (bovine CyP40, PDB 1iip) and FKBP (Arabidopsis thaliana FKBP42, PDB 2if4) sequences, are so marked (in color: Green = β-strand; Red = α-helix; Blue = unstructured coiled-coil) (Supplementary Material 3). Bottom: Rosetta ab initio prediction of disordered region is shown below, along with the corresponding amino acid sequence, which is also located within the linker. (B) Plots for seven in silico constructs with the same linker region positioned every 40 amino acids. The constructs are code-named by linker location, e.g., N40 means that the linker starts after 40 amino acids from the N-terminus (residue 41 to 92). (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
Disorder plot for Treponema denticola CFBP with relocated linker sequences. Disorder analysis and plot were performed essentially as in the two preceding Figures (Fig. 4, Fig. 5). (A) Top: Plot for native T. denticola CFBP with the 52-amino acid long linker region in its natural location. In addition, secondary structural elements, corresponding to known structures of homologous CYN (bovine CyP40, PDB 1iip) and FKBP (Arabidopsis thaliana FKBP42, PDB 2if4) sequences, are so marked (in color: Green = β-strand; Red = α-helix; Blue = unstructured coiled-coil) (Supplementary Material 3). Bottom: Rosetta ab initio prediction of disordered region is shown below, along with the corresponding amino acid sequence, which is also located within the linker. (B) Plots for seven in silico constructs with the same linker region positioned every 40 amino acids. The constructs are code-named by linker location, e.g., N40 means that the linker starts after 40 amino acids from the N-terminus (residue 41 to 92).