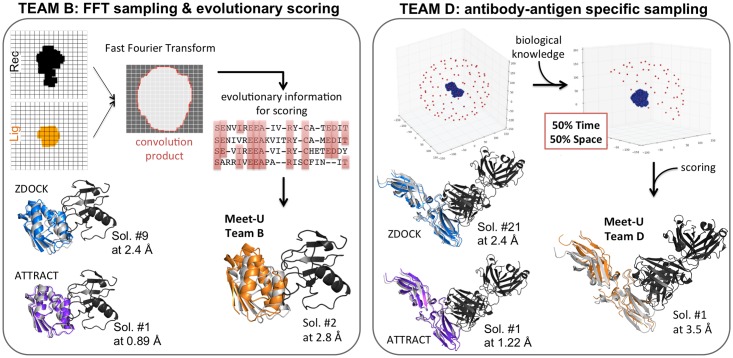
Fig 2. Examples of strategies and results for the 2016–2017 edition.
Left panel: Team B implemented an efficient sampling algorithm using a grid representation of the proteins to be docked and FFT. For the scoring, they used evolutionary information extracted from multiple sequence alignments of homologs of the two partners. Right panel: Team D used biological knowledge during the sampling step to filter out conformations early and drastically reduce the search space. The results obtained by the students (Teams B and D) on two complexes (barnase–barstar complex, Protein Data Bank [PDB] code: 1AY7, and an antibody–antigen complex, PDB code: 1JPS, respectively) are comparable to those obtained from state-of-the-art methods, namely ZDOCK [10] and ATTRACT [11]. ZDOCK relies on efficient sampling using FFT and on an optimized energy function [10]. ATTRACT proceeds through minimization steps using an empirical, coarse-grained molecular mechanics potential [11]. Candidate conformations for the complexes are represented as cartoons and superimposed onto the known crystallographic structures. The receptor is in black, the ligand from the candidate conformation is colored (in orange for Meet-U students, blue for ZDOCK, and purple for ATTRACT), and that from the crystallographic structure is in grey. With each candidate conformation are associated its rank, according to the scoring function of the method, and its deviation (in Å) from the crystallographic structure. FFT, Fast Fourier Transform; PDB, protein data bank.