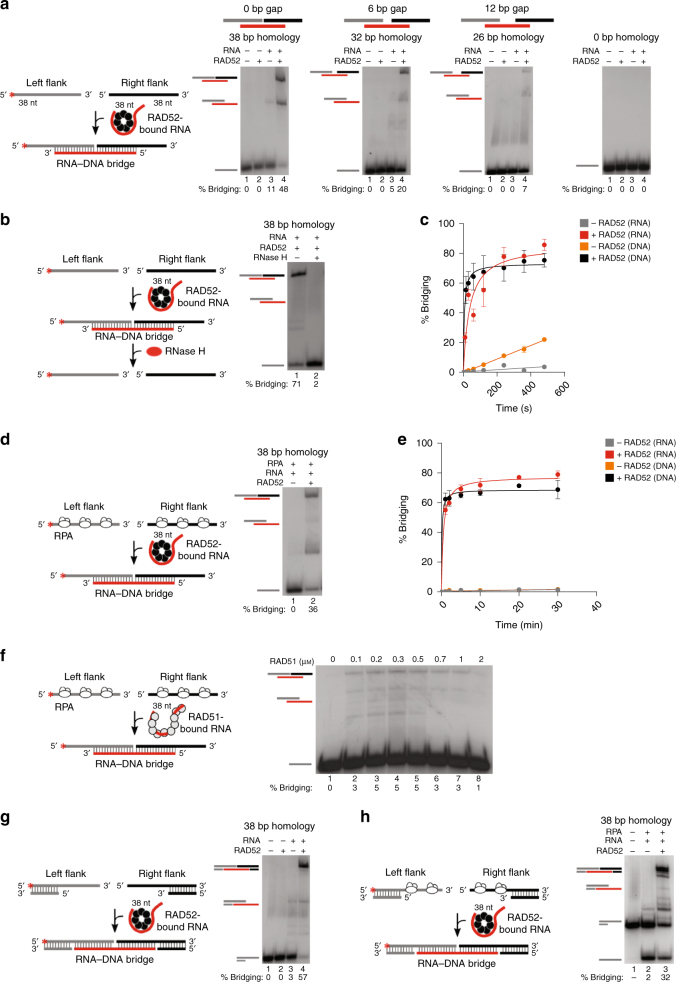
Fig. 2.
RAD52 promotes RNA-dependent DNA recombination. a Schematic of assay (left). Non-denaturing gels showing RAD52 RNA−DNA recombination (RNA-bridging of homologous DNA) in the presence of the indicated substrates (right). b Schematic of assay (left). Non-denaturing gel showing RNase H digestion of a RAD52-mediated RNA−DNA recombination intermediate (RNA−DNA recombinant bridge) (right). c Graph showing a time course of RNA–DNA recombination (bridging) compared to DNA−DNA recombination (bridging) of left and right flanking ssDNA without RPA and in the presence and absence of RAD52. Data shown as average ± SD, n = 3. d Schematic of assay (left). Non-denaturing gel showing RAD52 RNA−DNA recombination in the presence of the indicated RPA-coated substrates (right). e Graph showing a time course of RNA−DNA recombination (bridging) compared to DNA−DNA recombination (bridging) of left and right flanking RPA-bound ssDNA in the presence and absence of RAD52. Data shown as average ± SD, n = 3. f Schematic of assay (left). Non-denaturing gel showing RAD51 RNA−DNA recombination (bridging) in the presence of RPA pre-coated substrates (right). g Schematic of assay (left). Non-denaturing gel showing RAD52 RNA−DNA recombination (bridging) of the indicated pssDNA substrates (right). h Schematic of assay (left). Non-denaturing gel showing RAD52 RNA−DNA recombination (bridging) of the indicated RPA-coated pssDNA substrates (right). * = 32P label. % bridging indicated