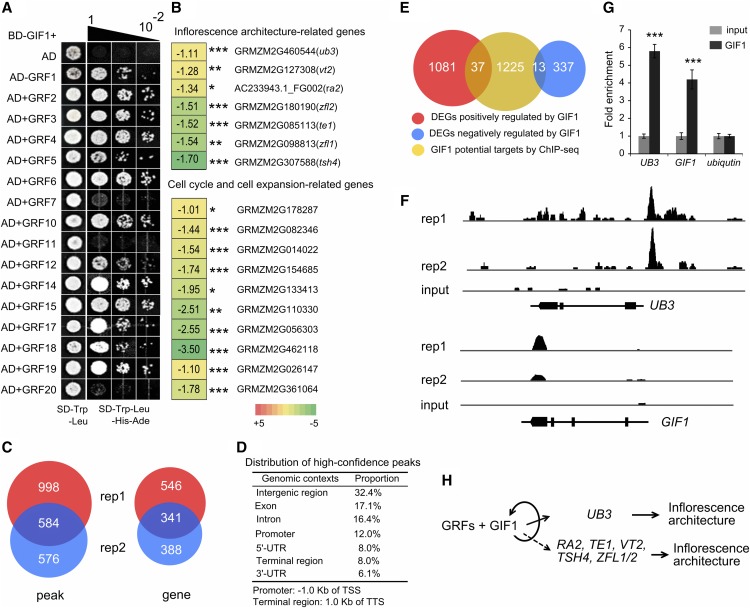
Figure 6.
GIF1-Interacting Proteins and Association Mapping.
(A) Identification of GIF1-interacting GRFs by yeast two-hybrid analysis. BD, binding domain; AD, activation domain.
(B) Expression changes for inflorescence architecture genes, and cell cycle and cell expansion genes in ∼5-mm tassels of gif1-1 compared with wild-type siblings. The number in the box represents expression alteration of the gene in log2 (RPKM in gif1-1/RPKM in the wild type). *P < 0.001, **P <0.0001, and ***P <0.00001.
(C) Venn diagram showing the number of peaks and genes identified in two biological replicates. Each replicate contained 10 tassels. rep, replicate.
(D) Distribution of high-confidence peaks. GIF1 binds in various genomic contexts, with high proportion of peaks located within genes. TSS, transcription start site; TTS, transcription termination site.
(E) Venn diagram representing a comparison between RNA-seq and ChIP-seq results.
(F) GIF1-bound regions in UB3 and GIF1. GIF1 can binds to its own fourth exon and 3′-UTR.
(G) PCR verification of GIF1-bound regions. The columns are the mean value of fold enrichment detected in three separate experiments, each with three technical replicates. Error bar show the sd. The statistical significance was estimated using a Student’s t test. *P < 0.05, **P < 0.01, and ***P < 0.001.
(H) A hypothetical model describing the GIF1 regulatory pathway in maize inflorescence development.