Fig. 3.
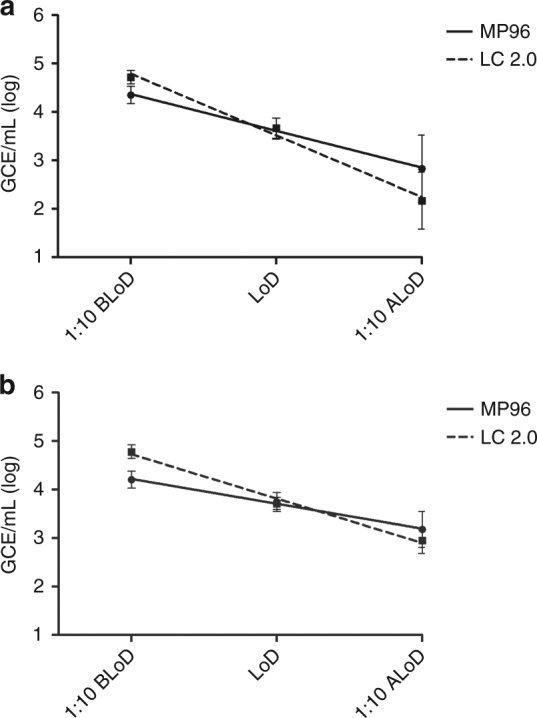
Analytical performance comparison between MP96 and LC 2.0 RNA extraction platforms in whole blood (EDTA). A pool of whole blood donated by healthy donors was contrived with ZIKV at a dilution of 1:10 before the limit of detection (1:10 BLoD), at the limit of detection (LoD) and at 1:10 after the limit of detection (1:10 ALoD). Twenty replicas of every dilution were extracted using small volume protocol (0.2 mL) and tested with Trioplex assay on the ABI7500 Fast Dx instrument (a) or in the QuantStudio Dx instrument (b). Mean genome copy equivalents per milliliter (GCE/mL) of viral RNA detected are displayed at each dilution. Error bars represent GCE/mL standard deviation. Straight line represents linear regression of MP96 platform (Roche). Dashed line represents linear regression of LC 2.0 platform (Roche)
