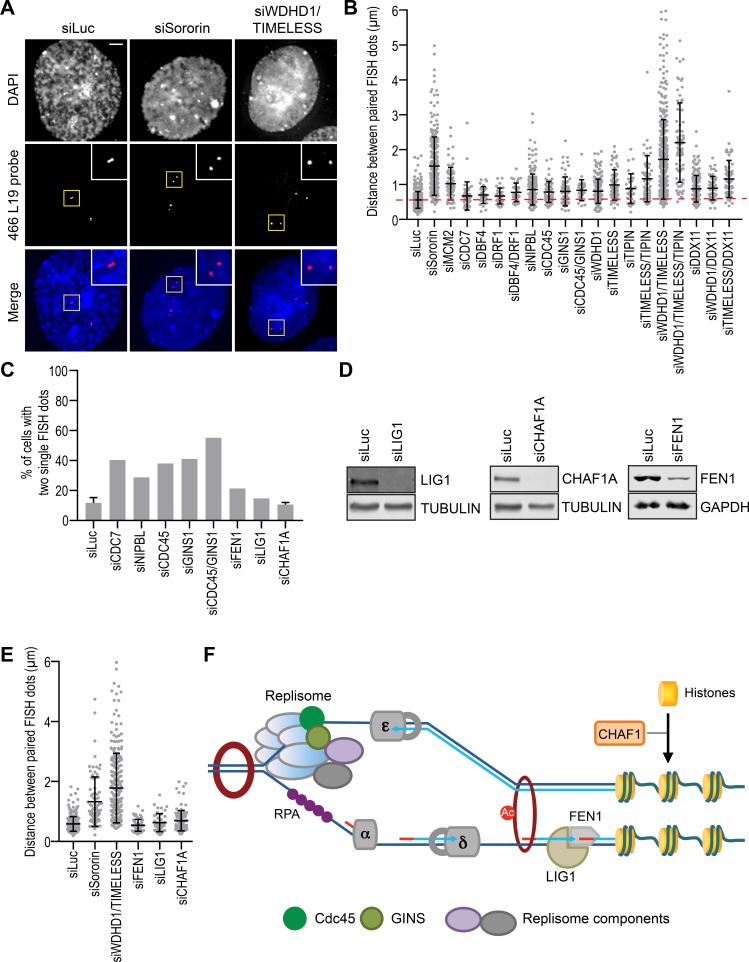
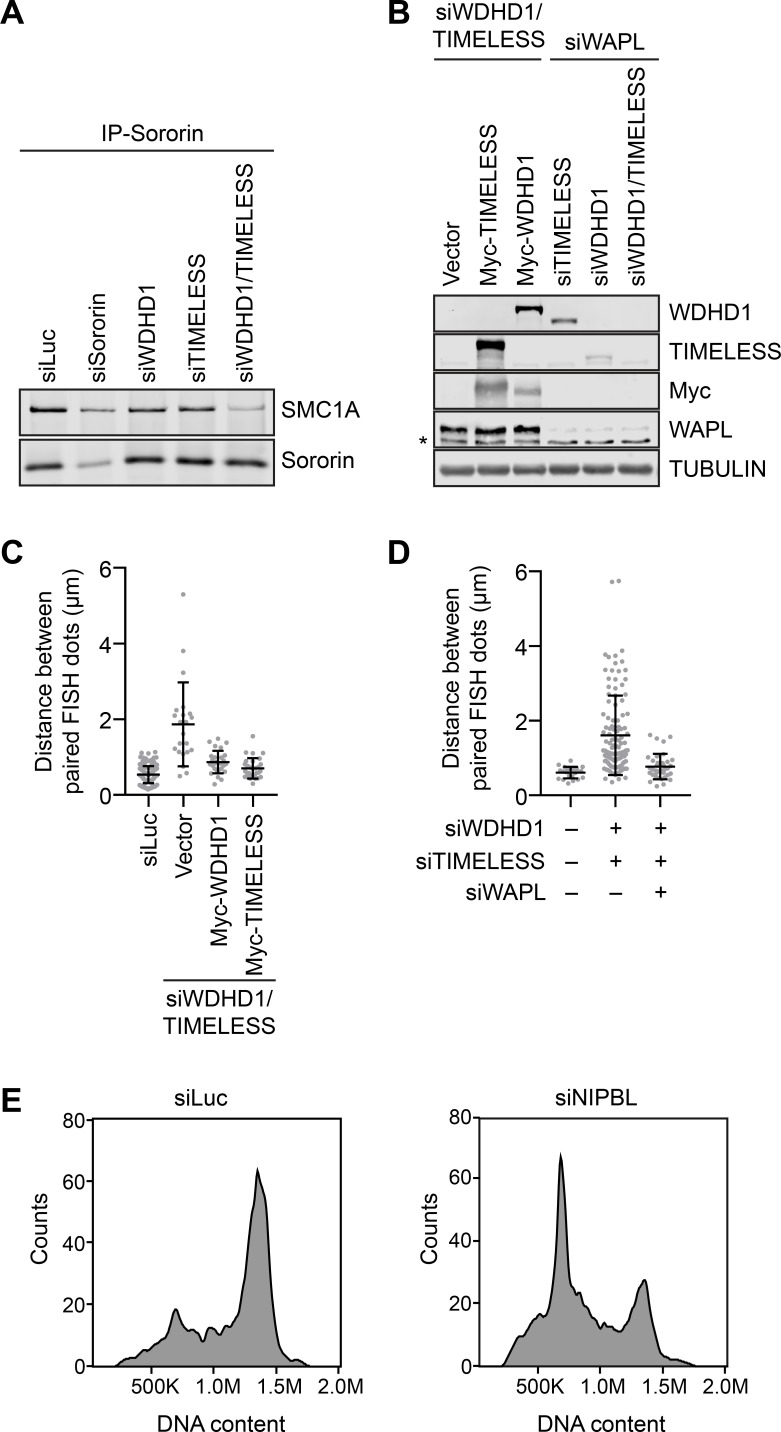
Figure 4. Replisome components are required for interphase sister-chromatid cohesion.
(A) Representative images of G2-enriched HeLa cells transfected with the indicated siRNAs and stained with DAPI (blue in merge) and the FISH probe (red in merge). Cells were treated with thymidine for 16–18 hr and released into fresh medium for 4 hr before fixation. Selected paired FISH signals are magnified in inset. Scale bar, 5 μm. (B) Quantification of the distances between paired FISH signals in G2-enriched HeLa cells transfected with the indicated siRNAs. Mean ± SD (siLuc, n = 502; siSororin, n = 243; siMCM2, n = 70; siCDC7, n = 44; siDBF4, n = 30; siDRF1, n = 30; siDBF4/DRF1, n = 32; siNIPBL, n = 191; siCDC45, n = 66; siGINS1, n = 68; siCDC45/GINS1, n = 26; siWDHD1, n = 198; siTIMELESS, n = 58; siTIPIN, n = 31; siTIMELESS/TIPIN, n = 51; siWDHD1/TIMELESS, n = 375; siWDHD1/TIMELESS/TIPIN, n = 71; siDDX11, n = 154; siWDHD1/DDX11, n = 52; siTIMELESS/DDX11, n = 65). The red dashed line indicates the mean of the siLuc sample. (C) Quantification of the percentage of cells with two unreplicated single FISH dots. Cells were transfected with the indicated siRNAs, treated with thymidine for 16–18 hr, and released into fresh medium for 4 hr. siLuc, n = 507 (mean ± SD; four independent experiments); siCDC7, n = 109; siNIPBL, n = 66; siCDC45, n = 71; siGINS1, n = 139; siCDC45/GINS1, n = 116; siFEN1, n = 112; siLIG1, n = 115; siCHAF1A, n = 226 (mean ± SD; two independent experiments). (D) Lysates of HeLa cells transfected with the indicated siRNAs were blotted with the indicated antibodies. (E) Quantification of the distances between paired FISH signals in G2-enriched HeLa cells transfected with the indicated siRNAs. Mean ± SD (siLuc, n = 382; siSororin, n = 82; siWDHD1/TIMELESS, n = 226; siFEN1, n = 60; siLIG1, n = 44; siCHAF1A, n = 110). (F) A simplified model delineating the molecular events during DNA replication. Cohesion establishment can occur without the maturation of Okazaki fragments and the deposition of histones.