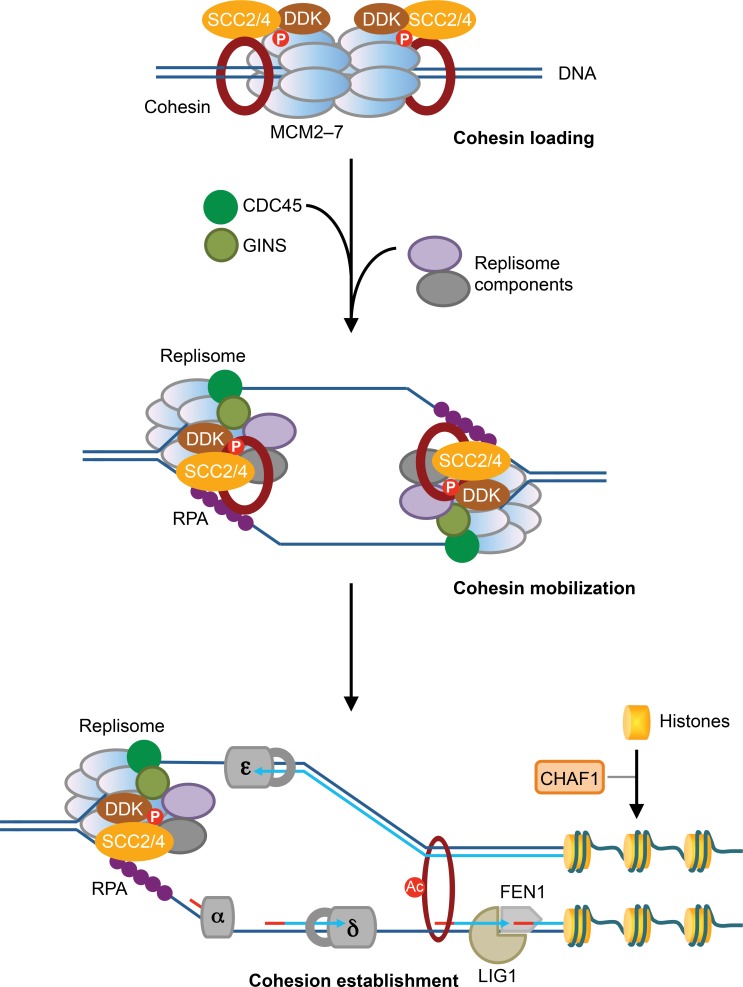
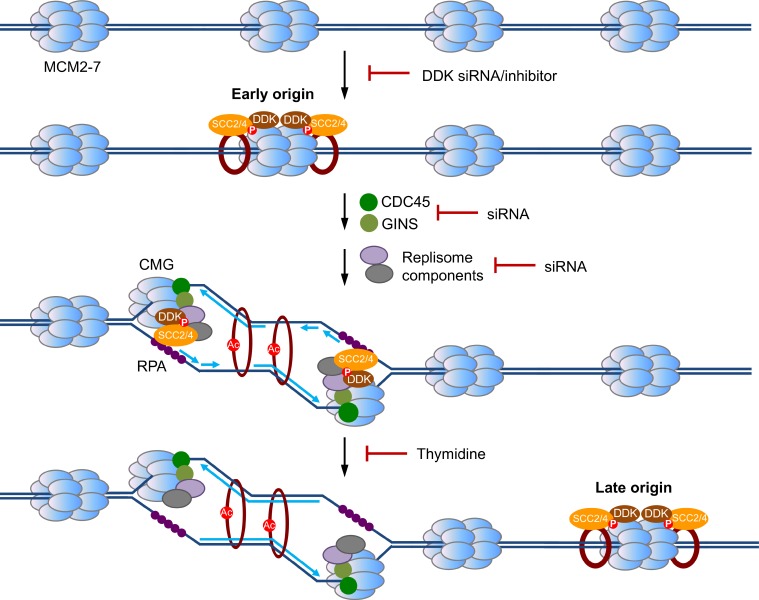
Figure 7. MCM2–7-mediated cohesin loading and mobilization promote sister-chromatid cohesion.
In this speculative model, SCC2/4 (NIPBL/MAU2 in humans) associates with DDK and the dormant, phosphorylated MCM2–7, and promotes cohesin loading at the G1/S boundary. The loaded cohesin remains physically associated with SCC2/4, DDK, and MCM2–7. Upon the activation of the helicase activity of MCM2–7 and the initiation of DNA replication, cohesin bound to the dormant MCM2–7 is mobilized and held at the active replication forks, through a process that requires SCC2/4, DDK, and a multitude of replisome components, including RPA. Cohesin bound to the replication fork is then deposited behind the fork prior to the completion of lagging strand synthesis and histone deposition, and entraps both sister chromatids to establish sister-chromatid cohesion.