Abstract
Background
Type III polyketide synthases (PKSs) are simple homodimer ketosynthases that distribute across plants, fungi, and bacteria, catalyzing formation of pyrone- and resorcinol-types aromatic polyketides with various bioactivities. The broad substrate promiscuity displayed by type III PKSs makes them wonderful candidates for expanding chemical diversity of polyketides.
Results
Violapyrone B (VLP B, 10), an α-pyrone compound produced by deepsea-derived Streptomyces somaliensis SCSIO ZH66, is encoded by a type III PKS VioA. We overexpressed VioA in three different hosts, including Streptomyces coelicolor M1146, Streptomyces sanyensis FMA as well as the native producer S. somaliensis SCSIO ZH66, leading to accumulation of different violapyrone compounds. Among them, S. coelicolor M1146 served as the host producing the most abundant violapyrones, from which five new (2–4, 7 and 12) and nine known (1, 5, 6, 8–11, 13 and 14) compounds were identified. Anti-influenza A (H1N1) virus activity of these compounds was then evaluated using ribavirin as a positive control (IC50 = 112.9 μM), revealing that compounds 11–14 showed considerable activity with IC50 values of 112.7, 26.9, 106.7 and 28.8 μM, respectively, which are significantly improved as compared to that of VLP B (10) (IC50 > 200 μM). The productions of 10 and 13 were increased by adding P450 inhibitor metyrapone. In addition, site-directed mutagenesis experiment led to demonstration of the residue S242 to be essential for the activity of VioA.
Conclusions
Biological background of the expression hosts is an important factor impacting on the encoding products of type III PKSs. By using S. coelicolor M1146 as cell factory, we were able to generate fourteen VLPs compounds. Anti-H1N1 activity assay suggested that the lipophilic nature of the alkyl chains of VLPs plays an important role for the activity, providing valuable guidance for further structural optimization of VLPs.
Electronic supplementary material
The online version of this article (10.1186/s12934-018-0908-9) contains supplementary material, which is available to authorized users.
Keywords: Violapyrones (VLPs), Overexpression, Type III polyketide synthase (PKS), Anti-influenza A (H1N1) virus activity
Background
Type III polyketide synthases (PKSs) catalyze carbon–carbon bond formation through a complete series of decarboxylation, condensation, and cyclization reactions with a single active site [1]. The diversity of type III PKS-catalyzed reactions is ascribed to selectivity of starter and extender units, number of condensations, and intramolecular cyclization manners [2]. Notably, many type III PKSs display broad substrate promiscuity, and can recognize unnatural substrates to generate novel unnatural products, thus making them fantastic candidates for enzymatic engineering to expand chemical diversity of polyketides [3–8].
Crystal structural studies have shown that type III PKSs share a common three-dimensional structure and catalytic machinery which contains a conserved Cys-His-Asn catalytic triad [9, 10]. Substitution of non-catalytic residues located at the substrate binding, CoA binding and cyclization pockets can change the preference of substrates, number of condensation as well as cyclization manner and thus have impacts on their product selectivity [11]. A lot of mutagenesis studies with the aim of broadening substrate specificity have been carried out in plant-derived type III PKSs. For example, the S338V variant of chalcone synthase (CHS) from Scutellaria baicalensis produced octaketides SEK4/SEK4b from eight molecules of malonyl-CoA instead of condensing 4-coumaroyl-CoA with three molecules of malonyl-CoA to generate naringenin chalcone as did the wild type CHS [12]. Substitution of N222 with Gly in octaketide synthase (OKS) from Aloe arborescens led to accumulation of a novel C20 decaketide SEK15 in addition to the C21 heptaketide chalcone that is produced by the wild type OKS [13]. The L214I variant of Vitis vinifera stilbene synthase (VvSTS) bears an increased substrate binding pocket and a decreased cyclization pocket compared with those in the wild type enzyme, resulting in production of short-chain polyketides with improved efficiency but absence of long-chain polyketides; conversely, the sizes of both pockets in the T197A variant were increased, thus leading to generation of five new polyketides which are not produced by the wild type VvSTS [14].
In contrast, only a handful of mutagenesis studies have been reported for prokaryotic type III PKSs, among which the active-site cavity-forming residue Y224 in Sg-RppA from Streptomyces griseus [15] and Sc-RppA from S. coelicolor [16] has been studied the most. In both enzymes, Y224 was demonstrated to be important for starter substrate selection, but Sc-RppA showed a higher tolerance towards certain amino acid changes of Y224 than Sg-RppA. Mutants of Y224 were thus generated, which preferentially recognize unnatural acyl-CoA such as acetyl-CoA, acetoacetyl-CoA, hexanoyl-CoA and benzoyl-CoA instead of malonyl-CoA as starter substrate [15, 16]. Another example is the mutagenesis of Gcs from S. coelicolor, which disclosed H261 and M274 are critical in controlling the substrate specificity and/or catalytic efficiency, as the H261A and M274A variants were capable of producing significantly increased amount of triketide pyrones in comparison to the wild-type Gcs [17].
Violapyrones (VLPs) are a group of α-pyrone compounds with antibacterial and anticancer activities [18–20]. Previously, we activated the VLP biosynthetic gene cluster via deletion of the global regulatory gene wblAso in deepsea-derived Streptomyces somaliensis SCSIO ZH66, leading to isolation of VLP B (10) (Fig. 1) [18]. This cluster is composed of a type III PKS gene vioA and a negative regulatory gene vioB. By inactivation of vioB, another four VLP compounds (VLP A, J, C and H) were obtained, which were subjected to evaluation of anti-MRSA (methicillin-resistant Staphylococcus aureus, MRSA) activity, demonstrating that the length of the alkyl side chains of VLPs played an essential role for the anti-MRSA activity [18].
Fig. 1.
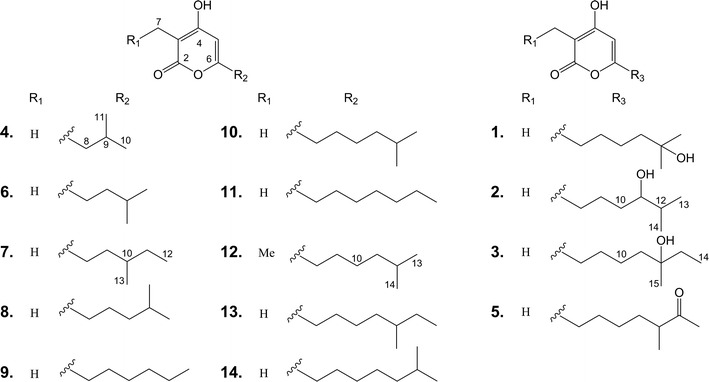
Structures of violapyrones (1–14). 1, VLP F; 5, VLP D; 6, VLP J1; 8, VLP A; 9, VLP J; 10, VLP B; 11, VLP I; 13, VLP C; 14, VLP H. 2, 3, 4, 7 and 12 are novel VLP analogues
The broad substrate promiscuity displayed by VioA inspired us to explore its synthetic potentials. Herein, we describe generation of VLP compounds (1–14) by over-expression of VioA in three different hosts, among which five (2–4, 7 and 12) are new. The following evaluation of anti-influenza A (H1N1) virus activity indicated that four (11–14) exhibited improved anti-H1N1 activity compared to that of VLP B (10).
Results
Phylogenetic analysis of VioA and distribution of vio cluster
To better understand the function of VioA, phylogenetic analysis was performed to with other characterized bacterial type III PKSs. As shown in Fig. 2, VioA belongs to the B2-2 clade [2], which preferentially uses short- and medium-chain (C2–C12) acyl-CoA as starter. VioA is closest to Cpz6 from the caprazamycin biosynthetic gene cluster [21], and they are clustered with DpyA and Gcs, which are proposed to recognize both CoA- and ACP-tethered β-keto acids from branched-chain or straight-chain fatty acid metabolism as starters, and to generate pyrones by lactonization of a linear polyketide intermediate [2]. In contrast, the members of the other subclade in B2-2 use malonyl CoA as both starter and extender unit to give scaffolds of pyrones and resorcinols by lactonization or Claisen-, aldol-type cyclization, respectively. We further mined the vioAB locus from other Streptomyces genomes in Genbank, and found another 21 Streptomyces genomes harboring vioAB homologous loci (Additional file 1: Table S3). Notably, most of them are located in a linear plasmid, probably contributing to their horizontal gene transfer during evolution.
Fig. 2.
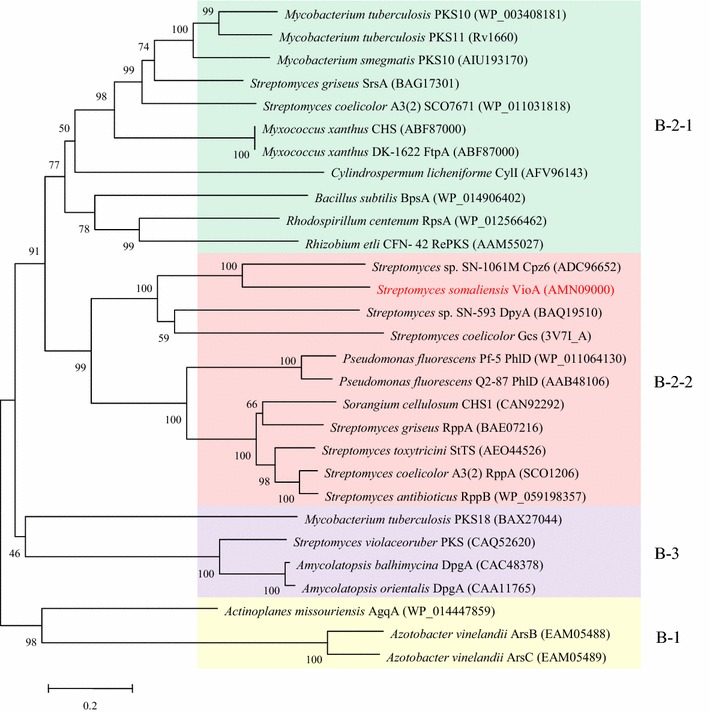
Phylogenetic analysis of VioA with characterized bacterial type III PKSs. Sequences were aligned with ClustalW, and the tree was constructed by using the neighbor joining method. The reliability of the tree was measured by bootstrap analysis with 1000 replicates. Scale: number of substitutions per nucleotide. Colored backgrounds indicate enzyme groups: yellow, B-1 PKS; green, B-2-1 PKS; pink, B-2-2 PKS; purple, B-3 PKS [2]
Overexpression of vioA in different Streptomyces strains
Type III PKSs capture acyl-CoA substrates from primary metabolism. Considering the variety of the acyl-CoA pools in different biological backgrounds, the VLP gene cluster was overexpressed in three different hosts, including the general heterologous expression host S. coelicolor M1146 [22], the marine-derived Streptomyces sanyensis FMA [23] as well as the native producer S. somaliensis SCSIO ZH66 [18, 24]. To get rid of the negative regulatory function of vioB [18], the vioA gene was put under the control of the constitutive promoter PgapDH followed by introduction into different hosts as described in the materials and methods section. HPLC analysis of the fermentation broths showed that in addition to VLP B (10), several other VLPs compounds were also accumulated in M1146/pWLI807 (Fig. 3i) and ZH66/pWLI807 (Fig. 3iii), conversely, only compound 1 was accumulated in FMA/pWLI807 (Fig. 3v), indicating the significant influence of the expression hosts on the products. Their relative yields in each host were indicated in Additional file 1: Figure S1.
Fig. 3.
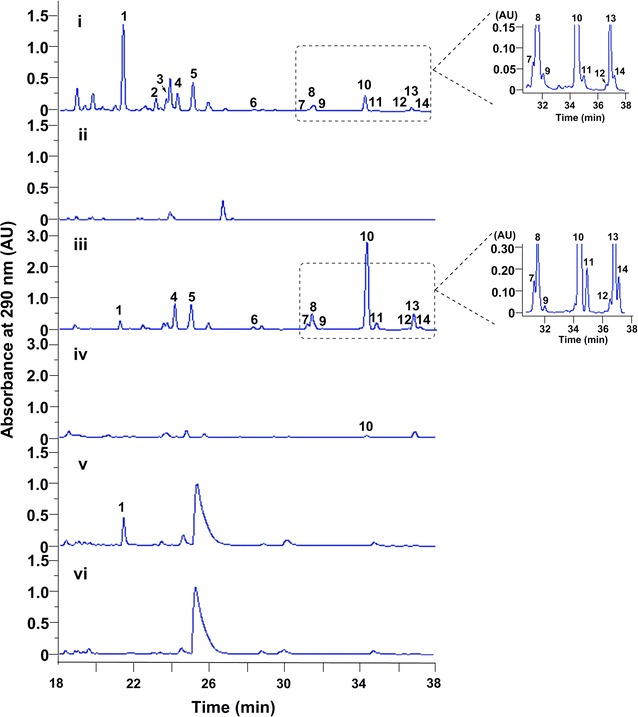
HPLC traces of the fermentation broths from overexpression strains. (i) S. coelicolor M1146/pWLI807; (ii) S. coelicolor M1146/pWLI806; (iii) S. somaliensis SCSIO ZH66/pWLI807; (iv) S. somaliensis SCSIO ZH66/pWL806; (v) S. sanyensis FMA/pWLI807; (vi) S. sanyensis FMA/pWLI806
Identification of the accumulated VLPs in the overexpression strains
From the large scale fermentations of the overexpression strain M1146/pWLI807, compounds 1–14 were isolated and identified via detailed NMR spectroscopic analysis.
Compound 2 was obtained as a colorless oil, and the molecular formula C13H20O4 was deduced from the [M+H]+ molecular ion peak at m/z 241.1432 in the HR-ESIMS (calcd for 241.1362). The structure of 2 was determined by 1D (1H and 13C) and 2D (COSY, HSQC and HMBC) NMR data. The 13C and HSQC spectra displayed three oxygenated quaternary carbons (δC 162.1–166.9), an olefinic methine carbon (δC 100.2), a quaternary carbon (δC 95.5), a hydroxylatedmethine carbon (δC 74.0), three methylene carbons (δC 32.7–33.2), a methine carbon (δC 23.2) and three methyl carbons (δC 8.3–18.9). Analysis of the COSY spectrum of 2 suggested a proton spin system from H-8 (δH 2.37) to H-13 (δH 0.81), constructing an aliphatic chain (Fig. 4). The location of the methyl group (δH 1.72) at C-3 (δC 95.5) was readily determined by its HMBC correlations (Fig. 4) with two oxygenated quaternary carbons C-2 (δC 165.4) and C-4 (δC 166.9), and as well as with C-3. Similarly, the olefinic methine proton H-5 (δH 5.93) showed HMBC correlations with C-3, C-4, C-6 (δC 162.1) and C-8 (δC 32.7) (Tables 1 and 2). From these HMBC correlations together with the fact that 2 needs to form a ring to satisfy the unsaturation number, an α-pyrone ring was constructed. In addition, the HMBC correlation from H-5 to C-8 confirmed the connectivity of the α-pyrone ring to the aliphatic chain. Thus, the structure of 2 was determined as 3-methyl-4-hydroxy-6-(11-hydroxy-12-methylhexyl)-2H-pyran-2-one, named as VLP L. The yield of 2 was 3.14 mg/L.
Fig. 4.
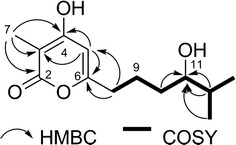
Key HMBC and COSY correlations of 2 in DMSO-d6
Table 1.
1H NMR (500 MHz) data for compounds 2–4, 7 and 12 in DMSO-d6 (J in Hz)
| Position | 2 | 3 | 4 | 7 | 12 |
|---|---|---|---|---|---|
| 2 | |||||
| 3 | |||||
| 4 | |||||
| 5 | 5.93, s, 1H | 5.94, s, 1H | 5.86, s, 1H | 5.89, s, 1H | 5.88, s, 1H |
| 6 | |||||
| 7 | 1.72, s, 3H | 1.72, s, 3H | 1.69, s, 3H | 1.70, s, 3H | 2.24, dd (14.5, 7.0), 2H |
| 8 | 2.37, m, 2H | 2.38, t (7.5), 2H | 2.21, d (6.5), 2H | 2.34, m, 2H | 2.35, t (7.0), 2H |
| 9 | 1.35, m, 2H | 1.49, m, 2H | 1.89, m, 1H | 1.52, m, 2H | 1.49, m, 2H |
| 10 | 1.27, m, 2H | 1.30, m, 2H | 0.88, d (6.5), 3H | 1.32, m, 1H | 1.26, m, 2H |
| 11 | 3.15, m, 1H | 1.30, m, 2H | 0.88, d (6.5), 3H | 1.13, m, 2H | 1.17, m, 2H |
| 12 | 1.50, m, 1H | 0.83, m, 3H | 1.50, m, 1H | ||
| 13 | 0.81, dd (10.0, 4.0), 3H | 1.32, m, 2H | 0.84, m, 3H | 0.84, d (6.5), 3H | |
| 14 | 0.81, dd (10.0, 4.0), 3H | 0.78, t (7.5), 3H | 0.84, d (6.5), 3H | ||
| 15 | 0.98, m, 3H | ||||
| 16 | 0.92, t (6.5), 3H |
Table 2.
13C NMR (125 MHz) data for compounds 2–4, 7 and 12 in DMSO-d6
| Position | 2 | 3 | 4 | 7 | 12 |
|---|---|---|---|---|---|
| 2 | 165.4 | 166.0 | 165.4 | 165.4 | 164.7 |
| 3 | 95.5 | 96.4 | 94.7 | 100.9 | 101.4 |
| 4 | 166.9 | 166.5 | 168.0 | 168.2 | 166.8 |
| 5 | 100.2 | 100.4 | 101.9 | 95.0 | 100.2 |
| 6 | 162.1 | 162.6 | 160.3 | 161.6 | 161.6 |
| 7 | 8.3 | 9.1 | 8.4 | 8.7 | 15.9 |
| 8 | 32.7 | 33.3 | 41.6 | 30.1 | 32.4 |
| 9 | 32.8 | 27.6 | 26.0 | 33.0 | 26.5 |
| 10 | 33.2 | 23.3 | 21.7 | 28.4 | 25.8 |
| 11 | 74.0 | 41.3 | 21.7 | 28.3 | 37.7 |
| 12 | 23.2 | 71.1 | 10.8 | 27.0 | |
| 13 | 18.9 | 34.5 | 18.5 | 22.2 | |
| 14 | 17.6 | 8.9 | 22.2 | ||
| 15 | 26.8 | ||||
| 16 | 12.5 |
Structure elucidation of compounds 3, 4, 7 and 12 was straightforward because of their close structural relationships with 2. Thus, by combination of NMR assignment with corresponding HR-ESIMS data, compounds 3, 4, 7 and 12 were identified as new VLP derivatives, named VLPs M-P, respectively. The 1H and13C chemical shift values of five new compounds (2–4, 7 and 12) are shown in Tables 1 and 2, respectively. The key HMBC and COSY correlations of 3, 4, 7 and 12 were described in supporting information (Additional file 1: Figures S4, S5, S8, S13). The yields of 3, 4, 7 and 12 were 1.75, 6.63, 0.29 and 0.13 mg/L, respectively.
Compounds 1, 5, 6, 8–11, 13 and 14 were identified as VLPs F, D, J1, A, J, B, I, C and H respectively, by comparison of 1H data with those reported in the literatures [18–20, 25] (Additional file 1: Figures S2, S6, S7, S9, S10, S11, S12, S14, S15), among which 8–10 and 13–14 had been identified from S. somaliensis SCSIO ZH66 mutant strains before [18]. Their yields of compounds 1, 5, 6, 8–11, 13 and 14 were 25.40, 7.50, 0.75, 4.73, 0.29, 8.53, 0.46, 2.97 and 0.25 mg/L, respectively.
Anti-H1N1 activity of VLPs
Before evaluating the anti-H1N1 activity of compounds 1–14, the cytotoxicity of compounds 1–14 in MDCK cell was evaluated by MTT assay [26]. The results in Table 3 showed that compounds 1–14 exhibited no significant cytotoxicity and CC50 value for compounds 1–14 were more than 1400 μM. Compounds 1–14 were evaluated for their anti-H1N1 activity by using CPE inhibition assay [27]. As shown in Table 3, compounds 11 and 13 showed moderate anti-H1N1 activities with IC50 values of 112.7 and 106.7 μM, respectively, which is comparable to that of the positive control ribavirin (IC50 = 112.9 μM); delightedly, compounds 12 and 14 exhibited stronger anti-H1N1 activities than ribavirin up to fourfold, with the IC50 values of 26.9 and 28.8 μM, respectively. In contrast, compounds 1–10 were inactive against H1N1 virus up to the concentration of 200 μM. Comparing the structures and bioactivities of these compounds, we proposed that the length and polarity of the alkyl side chains at C-3 and C-6 play essential roles for the antiviral activity, in which the one containing longer alkyl side chain with lower polarity gives better activity.
Table 3.
Inhibition effects of compounds 1–14 on H1N1 virus multiplication in vitro
| Compounds | IC50 (μM)a | CC50 (μM)b |
|---|---|---|
| 1–10 | > 200 | > 1500 |
| 11 | 112.7 | 2196.8 |
| 12 | 26.9 | 1565.5 |
| 13 | 106.7 | 1623.9 |
| 14 | 28.8 | 1451.7 |
| Ribavirin | 112.9 | 1517.5 |
The inhibition effects on influenza virus A/PR8/34 (H1N1) (MOI = 1.0) multiplication in MDCK cells were evaluated by virus yield reduction assay
aInhibition concentration 50% (IC50): concentration required to inhibit influenza virus A/PR8/34 (H1N1) yield at 48 h post-infection by 50%. Values are mean ± SD (n = 4)
bCytotoxic concentration 50% (CC50): concentration required to reduce MDCK cell viability by 50%. Values are mean ± SD (n = 3)
Site-directed mutagenesis of VioA
The broad substrate promiscuity of VioA makes itself an excellent candidate for further enzyme engineering to generate diverse VLPs. We next set out to further investigate the biosynthetic potentials of VioA in vitro. However, no soluble VioA was obtained after exploring different conditions (data not shown). Therefore, we turned to probe its function by expressing different versions of site-mutated vioA in S. coelicolor M1146. Firstly, we did multiple sequence alignment of VioA with selected reported type III PKSs (Additional file 1: Figure S16). With the purpose to probe the substrate promiscuity and/or to improve catalytic efficiency of VioA, I174 and L190 (corresponding to T197 and I214 in VvSTS, respectively), as well as Y229 and S242 (corresponding to H261 and M274 in Gcs, respectively) were substituted with Ala, Ile, Ala and Ala, respectively. Structure modeling was simultaneously performed as described in the Materials and Methods section to help understanding the underlying mechanism. As shown in Additional file 1: Figure S17A, both I174A (ii) and Y229A (iv) displayed severe defection on VLPs production, supporting their important roles in substrate binding (I174) and cyclization (Y229), respectively (Additional file 1: Figure S17B). No change was observed for L190I (Additional file 1: Figure S17A, iii), indicating this substitution probably had no influence on the cyclization pocket (Additional file 1: Figure S17B). Conversely, VLPs production was completely abolished in S242A (Additional file 1: Figure S17A, v), demonstrating S242 to be essential for the activity of VioA, which was consistent with its position being close to the cyclization pocket in the structural model (Additional file 1: Figure S17B).
P450 inhibitor increased the yields of anti-H1N1 VLPs by blocking side-chain oxidation
The above bioassay results indicated that the presence of hydroxyl- (1–3) or keto-group (5) in the alkyl side chain at C-6 has negative impact on the bioactivity. The introduction of hydroxyl- or keto-group might happen before (as an oxidized starter unit) or after the assembly of the pyrone ring (as a tailoring step). To test if they are assembled by cytochrome P450 monooxygenases, 2 mM of P450 inhibitor metyrapone was added into the fermentation medium. The results (Fig. 5) showed that the production of compounds 1–3 and 5 were decreased by ~ 2.3-, ~ 1.4-, ~ 3-, and ~ 7.8-fold, respectively, and simultaneously, the yields of 10 and 13 were increased by ~twofold and ~fourfold, respectively. Conversely, no obvious changes of the other compounds were observed. This result indicated that the presence of hydroxyl- or keto-group is assembled by an unknown P450 located in the genome of the heterologous host as proposed in Fig. 6.
Fig. 5.
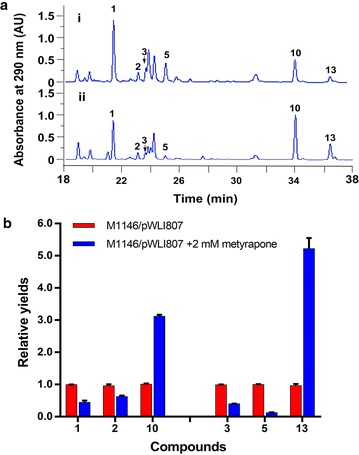
a HPLC traces of the fermentation broths from M1146/pWLI807 without metyrapone (i) and M1146/pWLI807 addition of 2 mM metyrapone (ii). b Relative yields of compounds 1–3, 5, 10 and 13 from M1146/pWLI807 without metyrapone and M1146/pWLI807 addition of 2 mM metyrapone
Fig. 6.
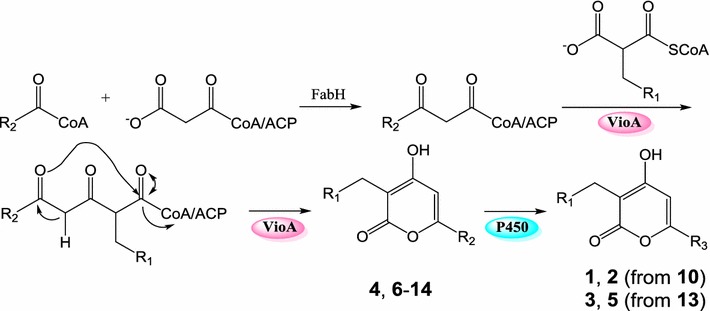
Proposed biosynthetic pathway of compounds 1–14
Discussion
Heterologous expression serves as a proven effective approach for activating silent secondary metabolites gene clusters [28–31]. As type III PKSs are simple homodimer ketosynthases, they are especially convenient to be manipulated. In this study, the type III PKS gene vioA from deepsea-derived S. somaliensis SCSIO ZH66 was put under the control of the constitutive promoter PgapDH followed by introduction into three different expression hosts. The accumulation of VLP products with different profiles in these three hosts (Fig. 3) supported the importance of precursor availability as well as genetic backgrounds of expression hosts.
The structural diversity of the VLPs compounds accumulated in S. coelicolor M1146/pWLI807 indicated that VioA can condense CoA- or ACP-tethered β-keto acids of different chain length with both ethylmalonyl CoA and methylmalonyl CoA, similar to that of Gcs [32]. However, compared to the data reported so far, VioA might recognize more diverse CoA- or ACP-tethered β-keto acids from fatty acid metabolism than Gcs. In the present study, although we tried to broaden the substrate promiscuity and/or improve catalytic efficiency of VioA via mutagenesis based on sequence alignment as well as previously mutagenesis results [14, 17], it is not surprising that no variants with expected properties were obtained. The VvSTS variants T197A and L214I were able to produce polyketides with different profiles than those of the wild-type VvSTS by changing the sizes of the substrate binding pocket and the cyclization pocket [14]; however, the corresponding substitutions in VioA led to severely impaired activity (I174A, Additional file 1: Figure S17A, ii) and no impact at all (L190I, Additional file 1: Figure S17A, iii). In Gcs, replacements of H261 and M274 with Ala significantly increased the yields of triketide pyrones [17]; on the contrary, the corresponding variants Y229A and S242A of VioA displayed severely impaired (Additional file 1: Figure S17A, iv) and totally abolished activity (Additional file 1: Figure S17A, v), respectively. The structure model of VioA supported the important roles of these mutated residues (Additional file 1: Figure S17B). Crystallographic studies would be contribute to disclose the underlying molecular basis for the substrate promiscuity of VioA and provide reliable guidance for further optimization.
Herein, for the first time, VLPs were shown to display anti-H1N1 virus activity (Table 3). Except for compound 12 harboring a 3-ethyl-4-hydroxy-α-pyrone ring, compounds 1–11, 13 and 14 all have a 3-methyl-4-hydroxy-α-pyrone backbone but with diverse side chains at C-6. The differences in their activity can be ascribed to the influence of the substituent at C-3 and the alkyl side chain at C-6. The anti-H1N1 activity increased with decrease in the polarity of the compounds, suggesting that the lipophilic nature of the alkyl chain plays an important role for the activity, which is consistent with their anti-MRSA assay results [18]. These findings indicated prospective directions for improving anti-H1N1 activity of VLPs.
Conclusion
The expression of vioA in S. coelicolor M1146 led to production of fourteen VLP compounds (1–14), among which five (2–4, 7 and 12) are novel compounds and four (11–14) display anti-H1N1 activities. Here, for the first time, VLPs were shown to display antiviral activity. The production of VLPs derivatives with enhanced antiviral activity were increased by adding P450 inhibitor-metyrapone. The site-directed mutagenesis results of VioA would provide reference for future enzyme engineering.
Methods
Strains and plasmids
All strains and plasmids used in this study are listed in Additional file 1: Table S1. Escherichia coli DH5α was used as the host for general subcloning [33]. E. coli ET12567/pUZ8002 [34] was used as the cosmid donor host for E. coli–Streptomyces intergeneric conjugation. The deepsea-derived S. somaliensis SCSIO ZH66 has been described previously [18, 24]. S. coelicolor M1146 [22] and S. sanyensis FMA [23] were used as the host strains for heterologous expression. Plasmid extractions and DNA purifications were carried out using standardized commercial kits (OMEGA, Bio-Tek, Guangzhou, China). PCR reactions were carried out with primers listed in Additional file 1: Table S2 using Pfu DNA polymerase (TIANGEN, Beijing, China). Oligonucleotide synthesis and DNA sequencing were performed by Sunny Biotech company (Shanghai, China). Restriction endonucleases and T4 DNA ligase were purchased from Fermentas (Shenzhen, China).
Growth conditions
Escherichia coli strains were routinely cultured in Luria–Bertani (LB) liquid medium at 37 °C, 200 rpm, or LB agar plate at 37 °C, with appropriate antibiotic selection. Streptomyces strains were grown at 30 °C on MS medium (3% soya flour, 2% mannitol, 2% agar powder) for sporulation and conjugation, and were cultured in Tryptic Soy Broth (TSB) medium (3% tryptic soya both, 10.3% sucrose, 0.1% tryptone, 0.05% yeast extract) for genomic DNA preparation. Fermentation medium consists of 1% soluble starch, 2% glucose, 4% corn syrup, 1% yeast extract, 0.3% beef extract, 0.05% MgSO4·7H2O, 0.05% KH2PO4, 0.2% CaCO3, and 3% sea salt, pH = 7.0.
Bioinformatic analysis
The evolutionary history was inferred using the Neighbor-Joining method [35]. The optimal tree with the sum of branch length = 10.17229024 is shown. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) are shown next to the branches. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Poisson correction method and are in the units of the number of amino acid substitutions per site. 29 protein sequences were used for analysis. All positions containing gaps and missing data were eliminated. There were a total of 301 positions in the final dataset. Evolutionary analyses were conducted in MEGA5 [36].
Overexpression of VioA
The vioA gene and the constitutive promoter PgapDH were amplified from the genome of S. somaliensis SCSIO ZH66 using primer pairs of vioAFP/vioARPBamHI and PgapDHFPEcoRI/PgapDHRP (Additional file 1: Table S2), respectively. After digestion with BamHI and EcoRI, the PCR products were cloned into the same sites of pMT3 and were confirmed by sequencing. The resulting plasmid pWLI902 was passed through E. coli ET12567/pUZ8002 and was then introduced into S. coelicolor M1146, S. sanyensis FMA and S. somaliensis SCSIO ZH66, respectively, via conjugation, according to the established procedures. Apramycin-resistant exconjugants were selected to afford S. coelicolor M1146/pWLI902, S. sanyensis FMA/pWLI902 and S. somaliensis SCSIO ZH66/pWLI902, respectively.
Production and analyses of VLPs
Spores of Streptomyces strains were inoculated into 50 mL of medium in a 250 mL flask for production analysis or into 200 mL in a 1 L flask for isolation, and were incubated at 30 °C, 220 r.p.m for 7 days. The culture supernatants were extracted twice with an equal volume of EtOAc. The combined EtOAc extracts were concentrated in vacuo to afford a brown residue, which was dissolved in MeOH, filtered through a 0.2 μm filter, and subjected to HPLC analysis. The HPLC system consisted of Agilent 1260 Infinity Quaternary pumps and a 1260 Infinity diode-array detector. Analytical HPLC was performed on an Eclipse C18 column (5 μm, 4.6 × 150 mm) developed with a linear gradient from 5% to 80% B/A in 40 min (phase A: 0.1% HCOOH in H2O; phase B: 100% ACN supplemented with 0.1% HCOOH) followed by an additional 10 min at 100% B at flow rate of 1 mL/min and UV detection at 290 nm. A total volume of 23 L fermentation cultures were harvested by centrifugation. The supernatant was treated as above and 3.95 g of brown residue was obtained, which was applied to reversed-phase C18 open column, eluting with a gradient eluent of H2O–MeOH (from 9:1 to 1:19 and finally 100% MeOH) to collect 17 fractions. Fraction 3 (366.21 mg) was further subjected to semipreparative HPLC using a YMC ODS-A column (250 × 20 mm i.d, 5 μm) by a linear gradient from 60 to 75% B/A in 80 min (phase A: 0.1% HCOOH in H2O; phase B: 100% MeOH supplemented with 0.1% HCOOH) to afford compound 1 (15.02 mg), compound 2 (5.74 mg), compound 3 (5.72 mg), compound 4 (1.70 mg) and compound 5 (5.05 mg). Compound 6 (3.97 mg) was obtained by further separation of fraction 5 (90.00 mg) eluting with 65% MeOH in H2O supplemented with 0.1% HCOOH. Compound 7 (3.20 mg), compound 8 (8.36 mg) and compound 9 (3.36 mg) were obtained by further separation of fraction 7 (53.37 mg) eluting with 75% MeOH in H2O supplemented with 0.1% HCOOH. Compound 10 (37.05 mg), compound 11 (3.88 mg) and compound 13 (2.58 mg) were purified from fraction 8 (152.11 mg) eluting with 80% MeOH in H2O supplemented with 0.1% HCOOH. Compound 12 (2.00 mg) and compound 14 (2.30 mg) were purified from fraction 9 (36.15 mg) by a linear gradient from 70% to 100% B/A in 60 min (phase A: 0.1% HCOOH in H2O; phase B: 100% MeOH supplemented with 0.1% HCOOH). The identities of VLPs were elucidated by HR-ESI-MS and NMR analysis. HR-ESI-MS was carried out on Thermo LTQ-XL mass spectrometer. NMR data was recorded with an Agilent-DD2500 spectrometer.
Site-directed mutation
For this work, the site-directed mutagenesis was created by overlapping primer mutagenesis [37]. To make each mutation, pairs of overlapping oligonucleotides, Additional file 1: Table S2, were synthesized. The first round of PCR was done using each of two mutagenic oligonucleotides and each of two (flanking) oligonucleotides complementary either to the 5′ or 3′ ends of the PgapDH::vioA. The two resulting PCR products were mixed, annealed and extended by few PCR cycles. The resulting Gel-purified full-length PCR products were cloned into pMT3 and confirmed by DNA sequencing.
Protein structure modeling
The structural model of VioA was done by using I-TASSER server (http://zhanglab.ccmb.med.umich.edu/I-TASSER) [38]. The C-score for the VioA model is 1.15, indicating a high degree of structural homology to the templates, which is additionally confirmed by the low RMSD of 4.1 ± 2.8 Å. COACH was then used for protein–ligand-binding site prediction [39, 40].
Biological assays
The cytotoxicity of compounds was measured by the MTT (3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyl tetrazolium bromide; Sigma–Aldrich, USA) assay. Confluent MDCK cell cultures in 96-well plates were exposed to different concentrations of compounds in triplicate, using incubation conditions equivalent to those used in the antiviral assays. Next, 10 μL of PBS containing MTT (final concentration: 0.5 mg/mL) was added to each well. After 4 h incubation at 37 °C, the supernatant was removed and 200 μL of DMSO was added to each well to solubilize the formazan crystals. After vigorous shaking, absorbance values were measured in a microplate reader (Bio-Rad, USA) at 570 nm. The CC50 was calculated as the compound concentration necessary to reduce cell viability by 50%. Compounds 1–14 were evaluated for their anti-influenza A (H1N1) virus activities by the cytopathic effect (CPE) inhibition assay [26]. Madin–Darby canine kidney (MDCK) cells were obtained from Cell Bank of Chinese Academy of Sciences (Shanghai, China) and grown in RPM1640 medium supplemented with 10% FBS, 100 units/mL of penicillin and 100 μg/mL of streptomycin. Influenza virus (A/Puerto Rico/8/34 [H1N1]; PR/8) was propagated in 10-day-old embryonated eggs for 3 days at 36.5 °C. MDCK cell cultures in 96-well plates were firstly infected with H1N1 virus (MOI = 0.1), and were then treated with different compounds in triplicate after removal of the virus inoculum. After 48 h incubation at 37 °C, the cells were fixed with 4% formaldehyde for 20 min at room temperature. After removal of the formaldehyde, the cells were stained with 0.1% crystal violet for 30 min. The plates were washed and dried, and the intensity of crystal violet staining for each well was measured at 570 nm. The virus inhibition (%) was calculated by the equation:
where Amock 570 was the absorbance without virus infection, Asample 570 was absorbance with virus infection and drug treatment, Avirus 570 was absorbance with virus infection but without drugs. Ribavirin injection (50 mg/mL) as a positive control was obtained from LuKang Cisen (China).
Additional file
Additional file 1: Table S1. Plasmids and strains used in this study. Table S2. Primer pairs used in this study. Table S3. Homologous locus of vioAB in different Streptomyces genomes. Figure S1. Relative yields for compounds 1–14 in different strains. Figure S2. Spectral data of 1. Figure S3. Spectral data of 2. Figure S4. Spectral data of 3. Figure S5. Spectral data of 4. Figure S6. Spectral data of 5. Figure S7. Spectral data of 6. Figure S8. Spectral data of 7. Figure S9. Spectral data of 8. Figure S10. Spectral data of 9. Figure S11. Spectral data of 10. Figure S12. Spectral data of 11. Figure S13. Spectral data of 12. Figure S14. Spectral data of 13. Figure S15. Spectral data of 14. Figure S16. Multiple-sequence alignments of VioA with selected type III PKSs. Figure S17. Site-directed mutagenesis study of VioA.
Authors’ contributions
LH and HH performed the experiments and wrote the draft manuscript. HL was involved in NMR analysis. SW assisted in biological assays. JJ sequenced the vio gene cluster. WL supervised the whole work and wrote the manuscript. All authors read and approved the final manuscript.
Acknowledgements
We would like to thank Dr. Lijiang Song from University of Warwick for helpful discussion, and thank Prof. Mervyn J. Bibb (John Innes Centre, UK) and Prof. Kui Hong (Wuhan University, China) for kindly providing us S. coelicolor M1146 and S. sanyensis FMA, respectively. This work was supported by Grants from the National Natural Science Foundation of China (31570032, 31711530219 and 31171201), and the NSFC-Shandong Joint Foundation (U1706206 and U1406402).
Competing interests
The authors declare that they have no competing interests.
Availability of data and materials
The datasets supporting the conclusions of this article are included within the article and its Additional files.
Ethics approval and consent to participate
Not applicable.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Abbreviations
- PKS
polyketide synthase
- VLP
violapyrone
- CHS
chalcone synthase
- OKS
octaketide synthase
- VvSTS
Vitis vinifera stilbene synthase
- Sg-RppA
Streptomyces griseus RppA
- Sc-RppA
Streptomyces coelicolor RppA
- Gcs
germicidin synthase
- wbl
whiB-like
- MRSA
methicillin-resistant Staphylococcus aureus
- Cpz6
caprazamycin 6
- DpyA
dihydroxyphenylglycine A
- ACP
acyl carrier protein
- NMR
nuclear magnetic resonance
- HPLC
high-pressure liquid chromatography
- HR-ESI-MS
high-resolution electrospray mass spectrometry
- COSY
correlation spectroscopy
- HSQC
heteronuclear singular quantum correlation
- HMBC
heteronuclear multiple bond correlation
- CPE
cytopathic effect
- IC50
half maximal inhibitory concentration
- UV
ultra violet
- MS
mannitol-soy flour
- LB
Luria–Bertani
- TSB
tryptic soy broth
- MDCK
Madin–Darby canine kidney
Footnotes
Electronic supplementary material
The online version of this article (10.1186/s12934-018-0908-9) contains supplementary material, which is available to authorized users.
Contributor Information
Lukuan Hou, Email: houlukuan1991@163.com.
Huiming Huang, Email: hmhuang1988@163.com.
Huayue Li, Email: lihuayue@ouc.edu.cn.
Shuyao Wang, Email: shuyaowang224@126.com.
Jianhua Ju, Email: jju@scsio.ac.cn.
Wenli Li, Email: liwenli@ouc.edu.cn.
References
- 1.Yu D, Xu F, Zeng J, Zhan J. Type III polyketide synthases in natural product biosynthesis. IUBMB Life. 2012;64(4):285–295. doi: 10.1002/iub.1005. [DOI] [PubMed] [Google Scholar]
- 2.Shimizu Y, Ogata H, Goto S. Type III polyketide synthases: functional classification and phylogenomics. ChemBioChem. 2017;18(1):50–65. doi: 10.1002/cbic.201600522. [DOI] [PubMed] [Google Scholar]
- 3.Zha W, Rubin-Pitel SB, Zhao H. Characterization of the substrate specificity of PhlD, a type III polyketide synthase from Pseudomonas fluorescens. J Biol Chem. 2006;281(42):32036–32047. doi: 10.1074/jbc.M606500200. [DOI] [PubMed] [Google Scholar]
- 4.Nakamura H, Hamer HA, Sirasani G, Balskus EP. Cylindrocyclophane biosynthesis involves functionalization of an unactivated carbon center. J Am Chem Soc. 2012;134(45):18518–18521. doi: 10.1021/ja308318p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Miyanaga A, Funa N, Awakawa T, Horinouchi S. Direct transfer of starter substrates from type I fatty acid synthase to type III polyketide synthases in phenolic lipid synthesis. Proc Natl Acad Sci. 2008;105(3):871–876. doi: 10.1073/pnas.0709819105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Wu HC, Li YS, Liu YC, Lyu SY, Wu CJ, Li TL. Chain elongation and cyclization in type III PKS DpgA. ChemBioChem. 2012;13(6):862–871. doi: 10.1002/cbic.201200051. [DOI] [PubMed] [Google Scholar]
- 7.Abe I. Novel applications of plant polyketide synthases. Curr Opin Chem Biol. 2012;16(1–2):179–185. doi: 10.1016/j.cbpa.2011.12.016. [DOI] [PubMed] [Google Scholar]
- 8.Morita H, Yamashita M, Shi SP, Wakimoto T, Kondo S, Kato R, Sugio S, Kohno T, Abe I. Synthesis of unnatural alkaloid scaffolds by exploiting plant polyketide synthase. Proc Natl Acad Sci USA. 2011;108(33):13504–13509. doi: 10.1073/pnas.1107782108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Abe I, Morita H. Structure and function of the chalcone synthase superfamily of plant type III polyketide synthases. Nat Prod Rep. 2010;27(6):809–838. doi: 10.1039/b909988n. [DOI] [PubMed] [Google Scholar]
- 10.Austin MB, Noel JP. The chalcone synthase superfamily of type III polyketide synthases. Nat Prod Rep. 2003;20(1):79–110. doi: 10.1039/b100917f. [DOI] [PubMed] [Google Scholar]
- 11.Bhan N, Li L, Cai C, Xu P, Linhardt RJ, Koffas MA. Enzymatic formation of a resorcylic acid by creating a structure-guided single-point mutation in stilbene synthase. Protein Sci. 2015;24(2):167–173. doi: 10.1002/pro.2600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Abe I, Watanabe T, Morita H, Kohno T, Noguchi H. Engineered biosynthesis of plant polyketides: manipulation of chalcone synthase. Org Lett. 2006;8(3):499–502. doi: 10.1021/ol052912h. [DOI] [PubMed] [Google Scholar]
- 13.Shi S-P, Wanibuchi K, Morita H, Endo K, Noguchi H, Abe I. Enzymatic formation of unnatural novel chalcone, stilbene, and benzophenone scaffolds by plant type iii polyketide synthase. Org Lett. 2009;11(3):551–554. doi: 10.1021/ol802606w. [DOI] [PubMed] [Google Scholar]
- 14.Bhan N, Cress BF, Linhardt RJ, Koffas M. Expanding the chemical space of polyketides through structure-guided mutagenesis of Vitis vinifera stilbene synthase. Biochimie. 2015;115:136–143. doi: 10.1016/j.biochi.2015.05.019. [DOI] [PubMed] [Google Scholar]
- 15.Funa N, Ohnishi Y, Ebizuka Y, Horinouchi S. Alteration of reaction and substrate specificity of a bacterial type III polyketide synthase by site-directed mutagenesis. Biochem J. 2002;367(3):781. doi: 10.1042/bj20020953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Li S, Gruschow S, Dordick JS, Sherman DH. Molecular analysis of the role of tyrosine 224 in the active site of Streptomyces coelicolor RppA, a bacterial type III polyketide synthase. J Biol Chem. 2007;282(17):12765–12772. doi: 10.1074/jbc.M700393200. [DOI] [PubMed] [Google Scholar]
- 17.Nguyen PHT. Probing substrate promiscuity of polyketide synthases. Master thesis. North Carolina State University; 2014.
- 18.Huang H, Hou L, Li H, Qiu Y, Ju J, Li W. Activation of a plasmid-situated type III PKS gene cluster by deletion of a wbl gene in deepsea-derived Streptomyces somaliensis SCSIO ZH66. Microb Cell Fact. 2016;15(1):116. doi: 10.1186/s12934-016-0515-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Shin HJ, Lee HS, Lee JS, Shin J, Lee MA, Lee HS, Lee YJ, Yun J, Kang JS. Violapyrones H and I, new cytotoxic compounds isolated from Streptomyces sp. associated with the marine starfish Acanthaster planci. Mar Drugs. 2014;12(6):3283–3291. doi: 10.3390/md12063283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zhang J, Jiang Y, Cao Y, Liu J, Zheng D, Chen X, Han L, Jiang C, Huang X. Violapyrones A-G, alpha-pyrone derivatives from Streptomyces violascens isolated from Hylobates hoolock feces. J Nat Prod. 2013;76(11):2126–2130. doi: 10.1021/np4003417. [DOI] [PubMed] [Google Scholar]
- 21.Kaysser L, Lutsch L, Siebenberg S, Wemakor E, Kammerer B, Gust B. Identification and manipulation of the caprazamycin gene cluster lead to new simplified liponucleoside antibiotics and give insights into the biosynthetic pathway. J Biol Chem. 2009;284(22):14987–14996. doi: 10.1074/jbc.M901258200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gomez-Escribano JP, Bibb MJ. Engineering Streptomyces coelicolor for heterologous expression of secondary metabolite gene clusters. Microb Biotechnol. 2011;4(2):207–215. doi: 10.1111/j.1751-7915.2010.00219.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Li T, Du Y, Cui Q, Zhang J, Zhu W, Hong K, Li W. Cloning, characterization and heterologous expression of the indolocarbazole biosynthetic gene cluster from marine-derived Streptomyces sanyensis FMA. Mar Drugs. 2013;11(2):466–488. doi: 10.3390/md11020466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zhang Y, Huang H, Xu S, Wang B, Ju J, Tan H, Li W. Activation and enhancement of Fredericamycin A production in deepsea-derived Streptomyces somaliensis SCSIO ZH66 by using ribosome engineering and response surface methodology. Microb Cell Fact. 2015;14:64. doi: 10.1186/s12934-015-0244-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yang RM, Zhang XL, Wang L, Huang JP, Yang J, Yan YJ, Luo JY, Wang XT, Huang SX. α-Pyrone derivatives from a streptomyces strain resensitize tamoxifen resistance in breast cancer cells. Nat Prod Bioprospect. 2017;7(4):329–334. doi: 10.1007/s13659-017-0136-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Senthilraja P, Kathiresan K. In vitro cytotoxicity MTT assay in Vero, HepG2 and MCF-7 cell lines study of marine yeast. J Appl Pharm Sci. 2015;5(3):080–084. doi: 10.7324/JAPS.2015.50313. [DOI] [Google Scholar]
- 27.Hung HC, Tseng CP, Yang JM, Ju YW, Tseng SN, Chen YF, Chao YS, Hsieh HP, Shih SR, Hsu JT. Aurintricarboxylic acid inhibits influenza virus neuraminidase. Antiviral Res. 2009;81(2):123–131. doi: 10.1016/j.antiviral.2008.10.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yamanaka K, Reynolds KA, Kersten RD, Ryan KS, Gonzalez DJ, Nizet V, Dorrestein PC, Moore BS. Direct cloning and refactoring of a silent lipopeptide biosynthetic gene cluster yields the antibiotic taromycin A. Proc Natl Acad Sci USA. 2014;111(5):1957–1962. doi: 10.1073/pnas.1319584111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Bilyk O, Sekurova ON, Zotchev SB, Luzhetskyy A. Cloning and heterologous expression of the grecocycline biosynthetic gene cluster. PLoS ONE. 2016;11(7):e0158682. doi: 10.1371/journal.pone.0158682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jordan PA, Moore BS. Biosynthetic pathway connects cryptic ribosomally synthesized posttranslationally modified peptide genes with pyrroloquinoline alkaloids. Cell Chem Biol. 2016;23(12):1504–1514. doi: 10.1016/j.chembiol.2016.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Nah HJ, Pyeon HR, Kang SH, Choi SS, Kim ES. Cloning and heterologous expression of a large-sized natural product biosynthetic gene cluster in Streptomyces species. Front Microbiol. 2017;8:394. doi: 10.3389/fmicb.2017.00394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Song L, Barona-Gomez F, Corre C, Xiang L, Udwary DW, Austin MB, Noel JP, Moore BS, Challis GL. Type III polyketide synthase β-ketoacyl-ACP starter unit and ethylmalonyl-CoA extender unit selectivity discovered by Streptomyces coelicolor genome mining. J Am Chem Soc. 2006;128(46):14754–14755. doi: 10.1021/ja065247w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: a laboratory manual. Cold Spring Harbor: Cold spring harbor laboratory press; 1989. [Google Scholar]
- 34.Paget MSB, Chamberlin L, Atrih A, Foster SJ, Buttner MJ. Evidence that the extracytoplasmic function sigma factor σE is required for normal cell wall structure in Streptomyces coelicolor A3(2) J Bacteriol. 1999;181(1):204–211. doi: 10.1128/jb.181.1.204-211.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4(4):406–425. doi: 10.1093/oxfordjournals.molbev.a040454. [DOI] [PubMed] [Google Scholar]
- 36.Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol. 2011;28(10):2731–2739. doi: 10.1093/molbev/msr121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Ho SN, Hunt HD, Horton RM, Pullen JK, Pease LR. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene. 1989;77(1):51–59. doi: 10.1016/0378-1119(89)90358-2. [DOI] [PubMed] [Google Scholar]
- 38.Yang J, Zhang Y. I-TASSER server: new development for protein structure and function predictions. Nucleic Acids Res. 2015;43(W1):W174–W181. doi: 10.1093/nar/gkv342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yang J, Roy A, Zhang Y. Protein-ligand binding site recognition using complementary binding-specific substructure comparison and sequence profile alignment. Bioinformatics. 2013;29(20):2588–2595. doi: 10.1093/bioinformatics/btt447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yang J, Roy A, Zhang Y. BioLiP: a semi-manually curated database for biologically relevant ligand–protein interactions. Nucleic Acids Res. 2013;41(Database issue):1096–1103. doi: 10.1093/nar/gks966. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: Table S1. Plasmids and strains used in this study. Table S2. Primer pairs used in this study. Table S3. Homologous locus of vioAB in different Streptomyces genomes. Figure S1. Relative yields for compounds 1–14 in different strains. Figure S2. Spectral data of 1. Figure S3. Spectral data of 2. Figure S4. Spectral data of 3. Figure S5. Spectral data of 4. Figure S6. Spectral data of 5. Figure S7. Spectral data of 6. Figure S8. Spectral data of 7. Figure S9. Spectral data of 8. Figure S10. Spectral data of 9. Figure S11. Spectral data of 10. Figure S12. Spectral data of 11. Figure S13. Spectral data of 12. Figure S14. Spectral data of 13. Figure S15. Spectral data of 14. Figure S16. Multiple-sequence alignments of VioA with selected type III PKSs. Figure S17. Site-directed mutagenesis study of VioA.
Data Availability Statement
The datasets supporting the conclusions of this article are included within the article and its Additional files.


