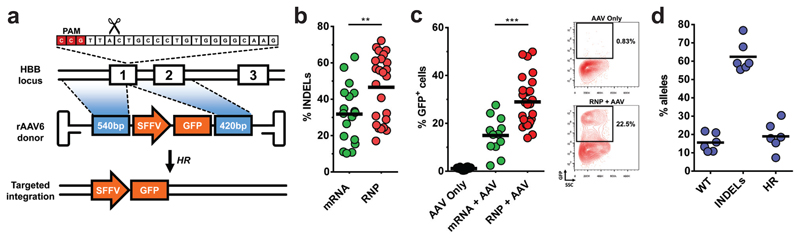
Fig 1. CRISPR/Cas9 and rAAV6-mediated targeted integration at the HBB locus in human CD34+ hematopoietic stem and progenitor cells (HSPCs).
a) Schematic of targeted genome editing at the HBB locus using CRISPR/Cas9 and rAAV6. Site-specific double strand breaks (DSBs) are created by Cas9 (scissors) mainly between nucleotide 17-18 of the 20bp target site, which is followed by the ‘NGG’ PAM (red). A DSB stimulates homologous recombination (HR) using rAAV6 homologous donor as repair template. White boxes: HBB exons, blue boxes: homology arms, orange boxes: SFFV-GFP-polyA expression cassette b) HSPCs were electroporated with all RNA or RNP CRISPR system and INDELs were analyzed via TIDE software (N=number of data points within group, all from different mPB or CB donors). c) HSPCs electroporated as above and transduced with HBB-specific rAAV6s were analyzed by flow cytometry 18-21 days post-electroporation when GFP levels were found to be constant. Left panel shows percentage of HSPCs. Right panel shows representative FACS plots (N=number of data points within group, all from different donors). d) HSPCs were treated as above but targeted with rAAV6 E6V donor. Frequencies of allele types were quantified by sequencing of a total of 600 clones from TOPO-cloned In-Out PCRs (N=6, all from different CB or BM donors)