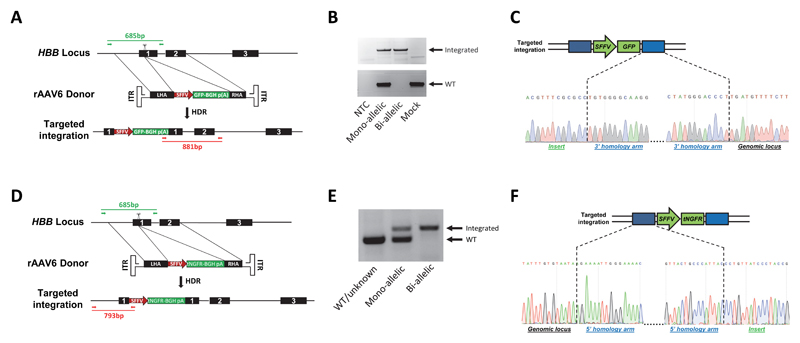
Extended Data Figure 4. Overview of PCR genotyping of methylcellulose colonies with HR of the GFP and tNGFR donor at the HBB locus.
(A) The HBB locus was targeted by creating a DSB in exon 1 via Cas9 (scissors) and supplying a rAAV6 GFP donor template. Alleles with integrations were identified by PCR (red, 881bp) on methylcellulose-derived colonies using an In-Out primer set. Wildtype alleles were identified by PCR (green, 685bp) using primers flanking the sgRNA target site. (B) Representative genotyping PCRs showing mono- and bi-allelic clones as well as a clone derived from Mock-treated cells. NTC = non-template control (see Suppl. Fig. 1a for uncropped gel). (C) Representative Sanger sequence chromatograms for junctions between right homology arm (in blue) and insert (in green) or genomic locus (in white) highlighting seamless homologous recombination. (D) The HBB locus was targeted by creating a DSB in exon 1 via Cas9 (scissors) and supplying a rAAV6 tNGFR donor template. Genotypes were assessed by a 3-primer genotyping PCR on methylcellulose-derived colonies using an In-Out primer set (red, 793bp) and a primer set flanking the sgRNA target site (green, 685bp). Note that the two forward primers are the same. (E) Representative genotyping PCRs showing a WT/unknown, mono-, and bi-allelic clone (see Suppl. Fig. 1b for uncropped gel). (F) Representative Sanger sequence chromatograms for junctions between left homology arm (in blue) and insert (in green) or genomic locus (in white) highlighting seamless homologous recombination.