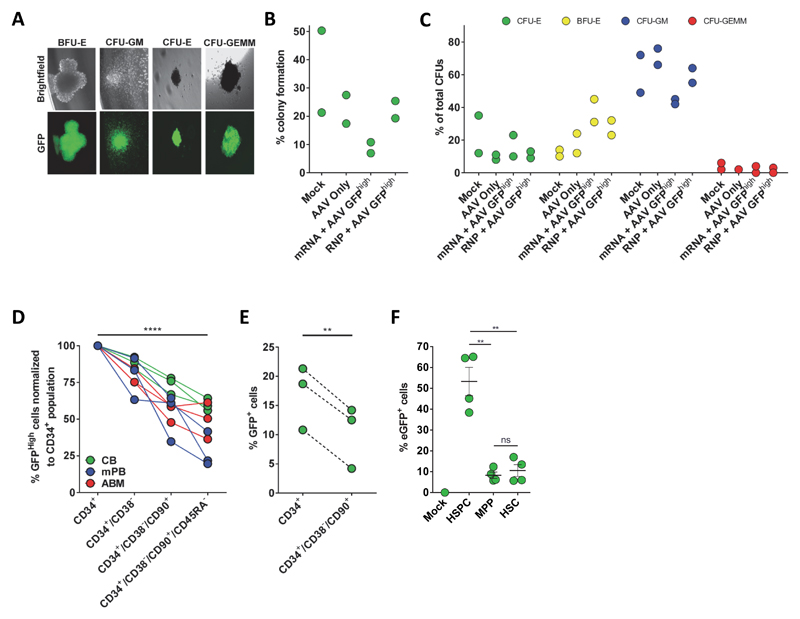
Extended Data Figure 5. Hematopoietic progenitor colony-forming unit (CFU) assay and targeting in different HSPC subpopulations.
(A) GFPhigh HSPCs were single cell-sorted into 96-well plates containing methylcellulose. Representative images from fluorescence microscopy show lineage-restricted progenitors (BFU-E, CFU-E, CFU-GM) and multipotent progenitors (CFU-GEMM) with GFP expression. (B) Colony forming units (CFUs) derived as described above were counted 14 days post sort and shown relative to the total number of cells sorted (% cloning efficiency) (N=2 different HSPC donors). (C) Colonies from above were scored according to their morphology: 1) CFU-Erythroid (CFU-E), 2) Burst Forming Unit-Erythroid (BFU-E), 3) CFU-Granulocyte/Macrophage (CFU-GM), and 4) CFU-Granulocyte/Erythrocyte/Macrophage/Megakaryocyte (CFU-GEMM). (N=2 different HSPC donors) (D) 500,000 HSPCs isolated from mobilized peripheral blood (mPB), adult bone marrow (ABM), or cord blood (CB) were electroporated with RNP and transduced with GFP rAAV6 donor. At Day 4 post-electroporation, cells were phenotyped by flow cytometry for the cell surface markers CD34, CD38, CD90, and CD45RA (Supplemental Figure 2). Percent GFPhigh cells in the indicated subpopulations are shown (data points represent unique donors, N=3 per HSPC source), **** p < 0.0001, paired Student’s t-test. (E) CD34+ or CD34+/CD38-/CD90+ cells were sorted directly from freshly isolated cord blood CD34+ HSPCs, cultured overnight, and then electroporated with RNP and transduced with GFP rAAV6. Bars show average percent GFP+ cells at Day 18 post-electroporation. Error bars represent S.E.M. (N=3 from different HSPC donors), ** p < 0.01, paired Student’s t-test. (F) MPP (CD34+/CD38-/CD90-/CD45RA-) and HSC (CD34+/CD38-/CD90+/CD45RA-) populations were sorted from fresh cord-blood-derived CD34+ HSPCs and immediately after sorting, cells were transduced with scAAV6-SFFV-eGFP at an MOI of 100,000 vg/cell along the bulk HSPC population. scAAV6 was used because it eliminates second strand synthesis as a confounder of actual transduction, though the activity of the SFFV promoter may not be equivalent in each population thus potentially underestimating the degree of transduction of MPPs and HSCs. Two days later, transduction efficiencies were measured by flow cytometric analysis of eGFP expression using non-transduced cells (Mock) to set the GFP+ gate. Error bars represent S.E.M., N=4, two different HSPC donors, ns = p ≥ 0.05, ** p < 0.01, unpaired t test with Welch's correction.