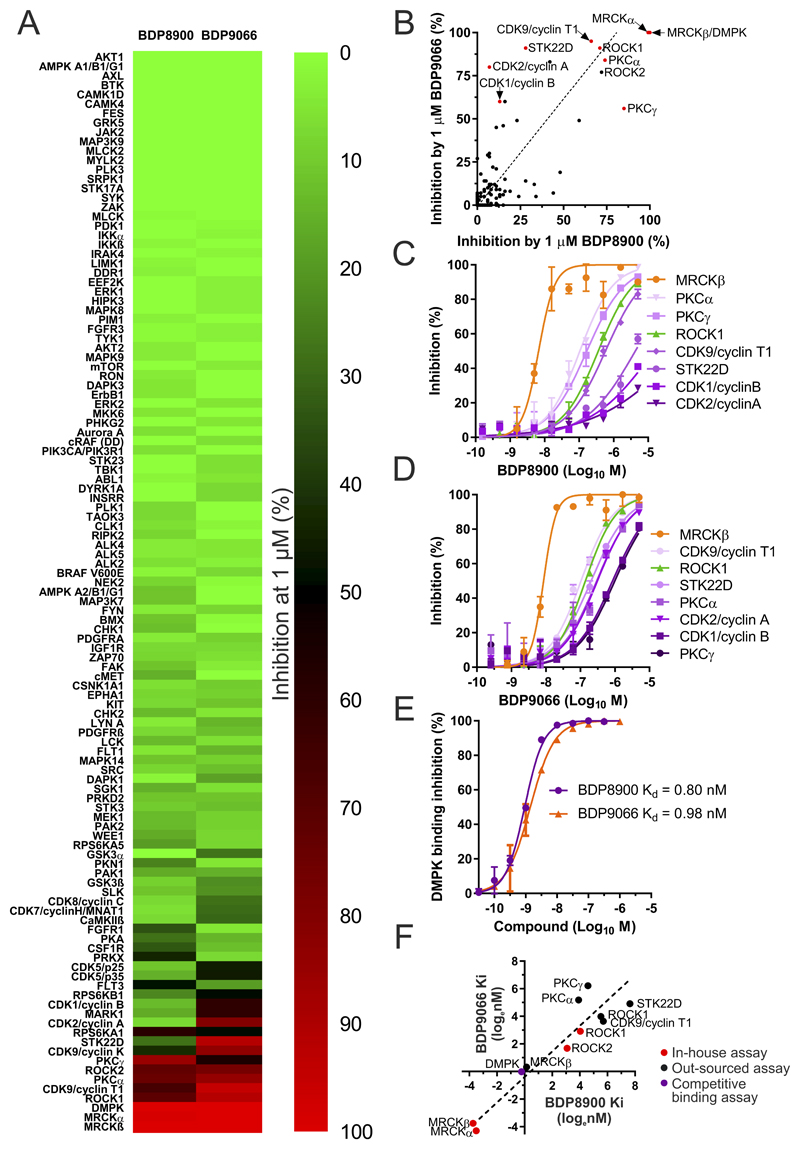
Figure 2. Detailed selectivity profiles for BDP8900 and BDP9066.
(A) Percentage kinase inhibition by 1 µM BDP8900 and BDP9066 were ranked and displayed by heat map. (B) Plot of inhibition by BDP8900 and BDP9066, kinases selected for detailed dose-response analysis indicated with red dot. ROCK1 was selected to additionally represent ROCK2 (black dot). Dotted line represents Deming regression (slope deviation from zero, p<0.0001). (C) BDP8900 dose-response curves for inhibition of indicated kinases in vitro. Results shown are mean ± SD of duplicate independent replicates. (D) BDP9066 dose-response curves for inhibition of indicated kinases in vitro. Results shown are mean ± SD of duplicate independent replicates. (E) BDP8900 and BDP9066 dose-response curves for competitive inhibition of tracer binding to DMPK in vitro, with calculated Kd values. Results shown are mean ± SD of duplicate independent replicates. (F) Natural log Ki or Kd (nM) values of BDP8099 and BDP9066 determined using in-house optimized assay conditions (red dots), by outsourced assays (black dots) or competition binding assay (purple dot). Dashed line represents Deming regression (slope deviation from zero, p<0.0001).