Fig. 7.
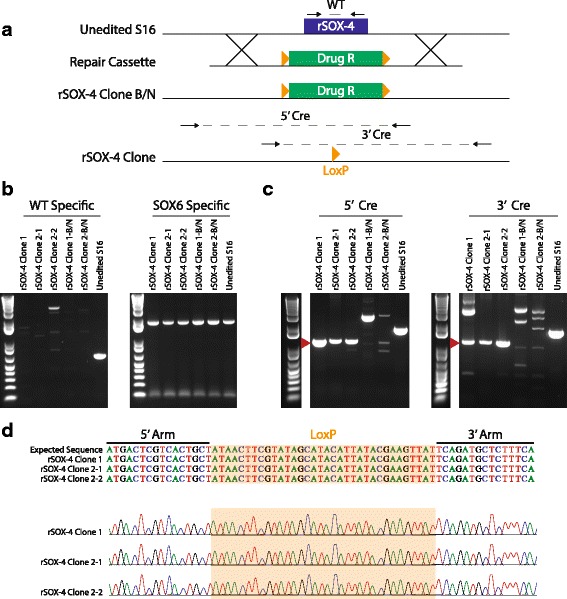
CRISPR-mediated deletion of rSOX-4 in S16 cells. a A cartoon depiction of the rSOX-4 deletion strategy is shown. The 651 bp rSOX-4 enhancer in unedited cultured rat Schwann (S16) cells is indicated by a blue rectangle. The drug resistance repair cassette is indicated by a green rectangle and the loxP sequences by orange triangles. Cross lines represent homologous regions for recombination-mediated repair during CRISPR/Cas9-mediated mutagenesis. Arrows represent the diagnostic PCR primers used in panels B and C, with the name of the corresponding PCR product [wild-type (WT), 5’ Cre, and 3’ Cre] above. b Wild-type (WT) specific PCR was performed on the three Cre:GFP-positive rSOX-4 clonal cell lines (Clone 1, Clone 2-1, Clone 2-2), the parental, pre Cre:GFP transfection (Clone1 – B/N and Clone 2 – B/N) cell lines, and unedited S16 cells. A SOX6-specific PCR was performed (right panel) as a DNA positive control. c Diagnostic PCR was performed across the 5′ (left panel) and 3′ (right panel) recombination sites for the same samples indicated in panel B using external (genomically anchored) and internal (repair template anchored) primers. The red arrowhead indicates the expected size for a single loxP scar. d Sequencing results from the rSOX-4 clonal cell lines in panel C (red arrowhead). The expected sequence was generated in silico based on proper recombination, Cre excision, and presence of a loxP scar
