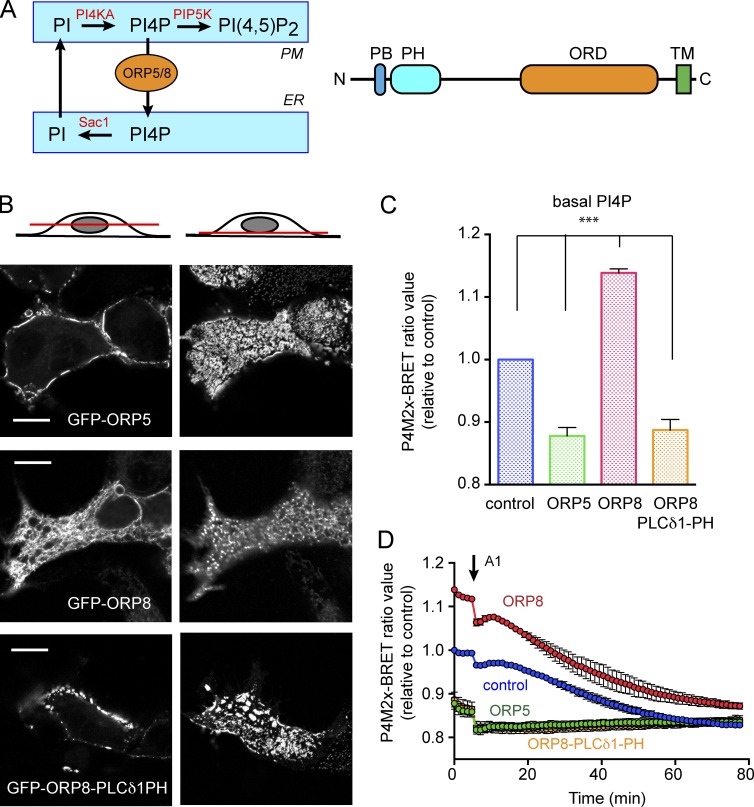
Figure 1.
PM interaction of ORP5/8 determines level of PI4P in the PM. (A) PI4P metabolism at ER-PM contact sites (left) and a linear domain structure of ORP5/8 (right). After being synthesized by PI4KA, PI4P is either phosphorylated by PIP5K to PI(4,5)P2 in the PM or transported to the ER. In the ER, PI4P is dephosphorylated by Sac1 phosphatase (left). Both ORP5 and ORP8 consist of a polybasic (PB) domain, a PH domain, an OSBP-related domain (ORD), and a transmembrane (TM) domain from N to C terminus (right). (B) Intracellular localization of ORP5, ORP8, and ORP8-PLCδ1-PH in live cells. HEK293-AT1 cells were transfected with GFP-tagged ORP5, ORP8, or ORP8-PLCδ1-PH and observed with confocal microscopy after 1 d. The same group of cells was observed on both the middle focal plane (left) and the bottom focal plane (right). Bars, 10 µm. (C) Quantitation of PM PI4P levels in live HEK293-AT1 cells with overexpression of ORP5, ORP8, or ORP8-PLCδ-PH by BRET analysis (see Materials and Methods for details). Grand means ± SEM are shown from three independent experiments performed in triplicate and normalized to the mean BRET value of mCherry-transfected (control) cells. Statistical significance was obtained with one-way ANOVA (P < 0.005). (D) Kinetics of PM-PI4P decrease by A1 treatment in BRET analysis. After measuring baseline BRET signal as shown in C, cells were treated with A1 (30 nM) and monitored for PI4P decrease. Grand means ± SEM are shown from three independent experiments performed in triplicate and normalized to the initial BRET value of mCherry-transfected (control) cells.