Figure 5.
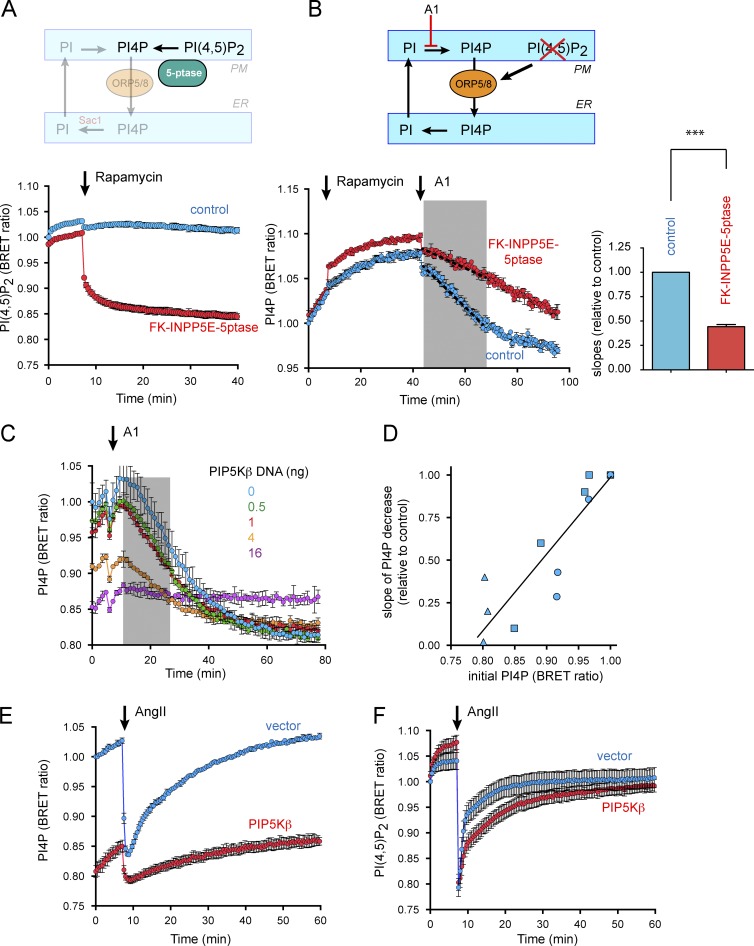
PI4P transport is controlled by PI(4,5)P2 levels in the PM. (A) Acute depletion of PM PI(4,5)P2 achieved by PM recruitment of INPP5E. The cartoon illustrates that PM-recruited 5ptase catalyzes dephosphorylation of PI(4,5)P2 to PI4P in the PM (top). Changes in PM PI(4,5)P2 levels were quantitated by BRET analysis before and after recruitment of 5ptase (bottom). HEK293-AT1 cells were cotransfected with mCherry-tagged FK-INPP5E and PM2-FRB before BRET analysis. Control cells were transfected with FK-PJ-dead plasmid instead of FK-INPP5E construct. After baseline measurement, cells were treated with DMSO or rapamycin for recruiting INPP5E to the PM. Relative PI(4,5)P2 levels were normalized to the initial BRET value of control cells. Grand means ± SEM are shown from three independent experiments performed in triplicate. (B) Rate of PI4P clearance controlled by PM PI(4,5)P2 levels. The cartoon illustrates that PM PI4P clearance after A1 treatment decreases when the ORP5/8-mediated transport is switched off as the 5ptase is recruited to the PM (top). PM PI4P clearance (after A1 addition) was monitored with or without PM PI(4,5)P2 depletion in BRET analysis (bottom). HEK293-AT1 cells were cotransfected with PM2-FRB and FK-INPP5E together with the BRET construct before BRET measurement. Control cells were transfected with FK-PJ-dead instead of FK-INPP5E. After baseline measurement, cells were treated with rapamycin for recruiting FK-PJ-dead (control) or -INPP5E to the PM and subsequently with A1 (30 nM) as indicated by the arrows. Grand means ± SEM are shown from three independent experiments performed in triplicate and normalized to the initial BRET value of control cells. The gray area shows the time period for which the rate of decline was calculated in each experiment. These rate values are shown in the column diagram relative to the rates calculated in the controls (***, P < 0.005; n = 3). (C) Representative BRET experiment showing rate of PI4P clearance from the PM under dose-dependent overexpression of myc-tagged PIP5Kβ. HEK293-AT1 cells were transfected with 0 ng (blue), 0.5 ng (green), 1 ng (red), 4 ng (orange), and 16 ng (purple) DNA per well of myc-PIP5Kβ before BRET analysis. Means ± SD are shown from triplicate experments, normalized to the initial BRET value of the cells with no myc-PIP5Kβ (blue). (D) Rate of PI4P clearance correlated with initial PM PI4P levels (square symbols). Two more BRET experiments were conducted by transfecting 0, 2, 6, and 12 ng PIP5Kβ DNA per well (circles) or 0, 4, 8, and 16 ng of the myc-PIP5Kβ DNA per well (triangles), also performed in triplicate. After treatment of A1, PI4P reduction rate (slope designated in y-axis) was calculated from the highest levels (initial PI4P designated in x-axis) for 1,000-s. Relative reduction rates were normalized to initial PI4P levels with no myc-PIP5Kβ in each experiment. (E) Effect of PIP5Kβ overexpression on PM PI4P levels responding to AngII stimulation. HEK293-AT1 cells were transfected with pcDNA3.1(HA) vector or myc-tagged PIP5Kβ. Change in PM PI4P levels was analyzed with BRET. After baseline measurement, cells were simulated with AngII (100 nM). Grand means ± SEM are shown from three independent experiments performed in triplicate and normalized to the initial BRET value of vector-transfected cells. (F) Effect of PIP5Kβ overexpression in PM PI(4,5)P2 levels responding to AngII stimulation. BRET analysis was conducted as described in E except that PM PI(4,5)P2 levels were monitored. Grand means ± SEM are shown from three independent experiments performed in triplicate and normalized to the initial BRET value of vector-transfected cells.