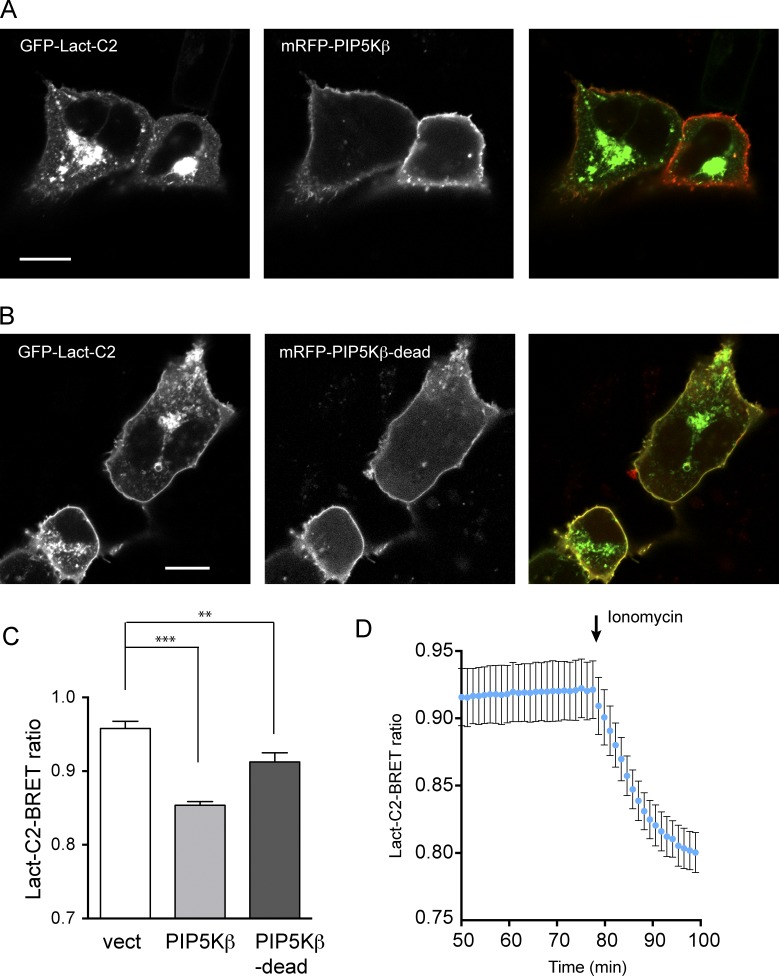
Figure 8.
PIP5Kβ-expressing cells show reduced PM PS levels. (A) Representative live-cell image displaying PS distribution under overexpression of wild-type PIP5Kβ. HEK293-AT1 cells were cotransfected with GFP-tagged Lact-C2 and mRFP-tagged PIP5Kβ and subjected to confocal microscopy 1 d after transfection. (B) Representative live-cell image displaying PS distribution under overexpression of kinase-dead PIP5Kβ. HEK293-AT1 cells were cotransfected with GFP-tagged Lact-C2 and mRFP-tagged PIP5Kβ-dead and subjected to confocal microscopy 1 d after transfection. Bars, 10 µm. (C) Quantitation of PM PS levels in wild-type or kinase-dead PIP5Kβ-expressing cells with BRET analysis. HEK293-AT1 cells were transfected with pcDNA3.1(HA) vector, myc-PIP5Kβ, or myc-PIP5Kβ-dead before BRET experiment. PM PS levels were analyzed by measuring mean emission intensity of PM-anchored Venus per Lact-C2-fused luciferase in the presence of coelenterazine h for 7 min. Grand means ± SEM are shown from three independent experiments performed in triplicate. Statistical significance was obtained with two-way ANOVA (***, P < 0.001; **, P < 0.005). (D) Dynamic range of PM PS levels quantitated with BRET. HEK293-AT1 cells were transfected with mCherry empty vector, and PM PS levels were analyzed by measuring mean intensity of PM-anchored Venus per Lact-C2-fused luciferase. After monitoring steady state under DMSO treatment, Ionomycin (10 µM) was treated to deplete PS in the inner leaflet of the PM resulting from PS externalization. Grand means ± SEM are shown from three independent experiments performed in triplicate.