FIGURE 2.
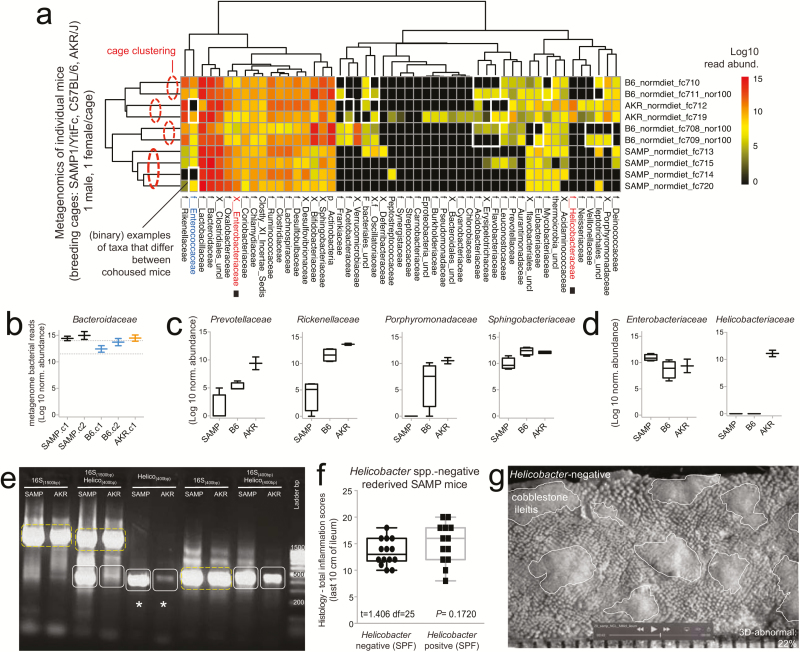
Metagenomic analysis reveals Helicobacter variability with minimal effect on SAMP ileitis. A, Individual fecal metagenomic analysis of breeder mice was performed 8 months after the previously pooled fecal sampling experiment, demonstrating bacterial profiles with reproducible cluster within pairs of breeders for each individual cage. Squares highlight absence of Helicobacteraceae in Helicobacteraceae-negative SAMP colony. When present, Helicobacteraceae was highly abundant in the feces of positive mice. Analysis highlights within-cage mouse microbiota individualities, eg, Enterococcaceae. B, Boxplots, showing averages for each cage, reveal high abundance of Bacteroidaceae in SAMP mice (compared with experiment 1 in Fig. 1D). C, The abundance of other Bacteroidetes is shown by boxplots. D, Compared with AKR and B6 mice, SAMP had a significantly higher level of Enterobacteracea (a family that contains Escherichia coli). Notice marked abundance of dichotomous variability of Helicobacteraceae in mice. E, Gel electrophoresis of PCR amplified target regions of Helicobacter spp. within the 16S rRNA gene from fecal DNA of random mice form our SAMP and AKR colony, 4 months after the individual fecal metagenomic experiment. Notice high band intensity in SAMP (asterisks). Universal and PCR-specific primers were used to generate full or partial gene amplicons (1500 or 400 bp). F, Histological inflammation scores of distal ileum from rederived Helicobacter-negative SAMP compared with that of the SAMP colony show no differences. Notice wider variability in Helicobacter-positive mice. G, Representative snapshot of videostereomicroscopy shows unchanged morphological appearance of ileal mucosal surface in Helicobacter-negative SAMP mice. Scale = 500 um.