Fig. 3.
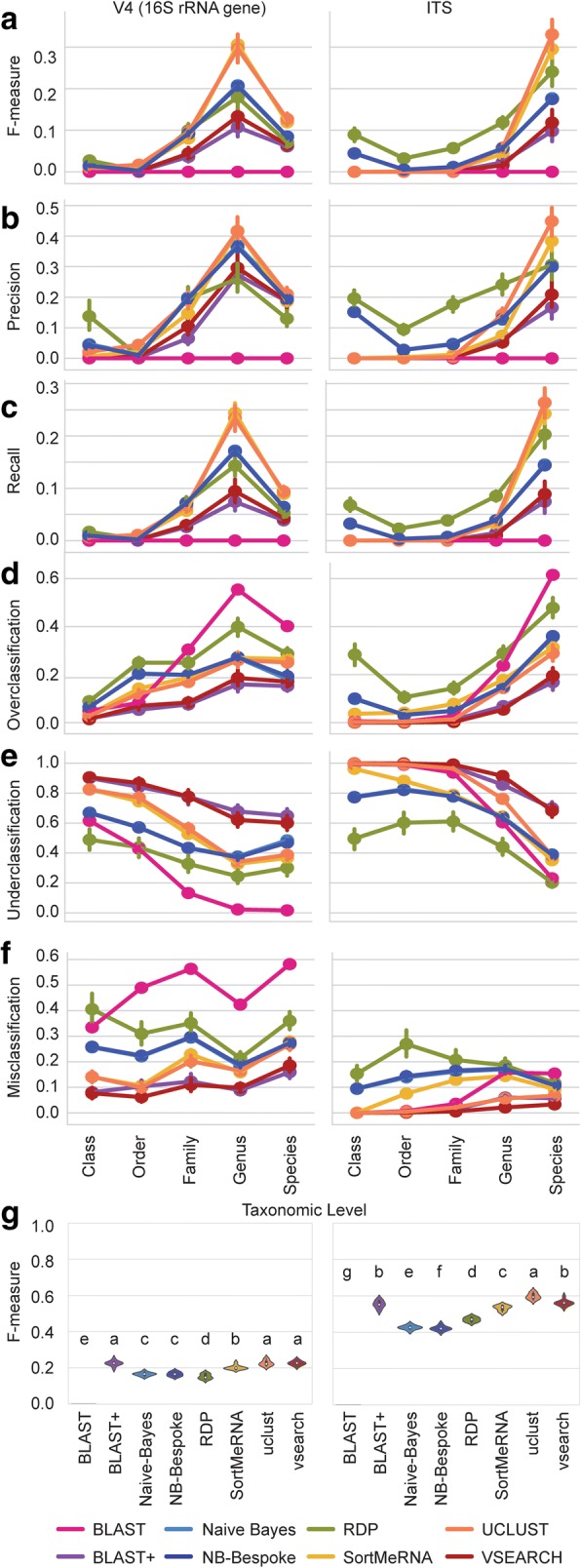
Classifier performance on novel-taxa simulated sequence datasets for 16S rRNA gene sequences (left column) and fungal ITS sequences (right column). a–f, Average F-measure (a), precision (b), recall (c), overclassification (d), underclassification (e), and misclassification (f) for each taxonomy classification method (averaged across all configurations and all novel taxa sequence datasets) from phylum to species level. Error bars = 95% confidence intervals. b Average F-measure for each optimized classifier (averaged across all novel taxa sequence datasets) at species level. Violins with different lower-case letters have significantly different means (paired t test false detection rate-corrected P < 0.05)
