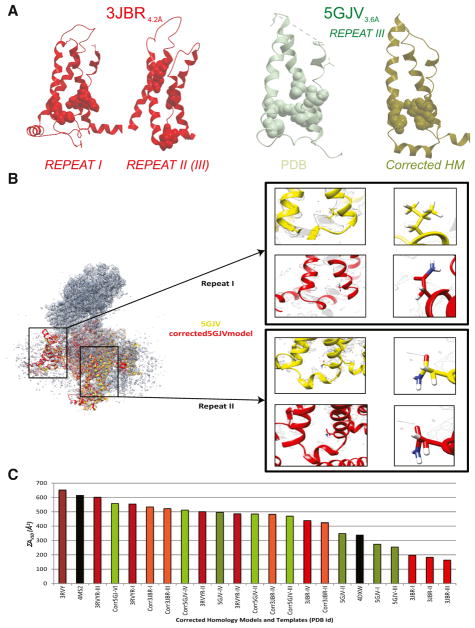
Figure 2. ΣAVSD Scores Correlate with Inaccuracies in Residue Assignment and Provide Structural Constraints for Accurate Homology Modeling of VGIC VSD.
(A–C) Ribbon, stick, and CPK representation of the VSD structural motif of PDB: 3JBR and PDB: 5GJV (A), both near-atomic cryo-EM structures of the Cav1.1 channel that lack the VSD interaction as a result of incorrect residue assignment of the functional residues. For PDB: 5GJV, we show that, by correctly placing the functional residues in their accurate locations, we reconstitute the VSD interaction in our corrected structure and produce a molecular model that is in higher agreement with the cryo-EM electron density map and raw data, as depicted (B) by the structural superimposition of the deposited 3D cryo-EM-based model (PDB: 5GJV) and our corrected-5GJV homology model, which shows that our model’s residue assignment correlates better with the cryo-EM density map than the deposited model. This is evidenced by the specific density pattern created by glutamate versus that of leucine, in which the oxygen atoms are expected to be distorted whereas the carbon atoms are expected to be resolved in the electron density map, as highlighted by the enlargements of the same amino acids in repeat I, which was incorrectly modeled, and repeat II, which was correctly modeled. Our corrected rabbit Cav1.1 model not only correlates better with the map, but also displays an improved ΣAVSD score over the models that have been deposited in the PDB, as shown in the graph (C). See also Figure S2.