Fig. 5.
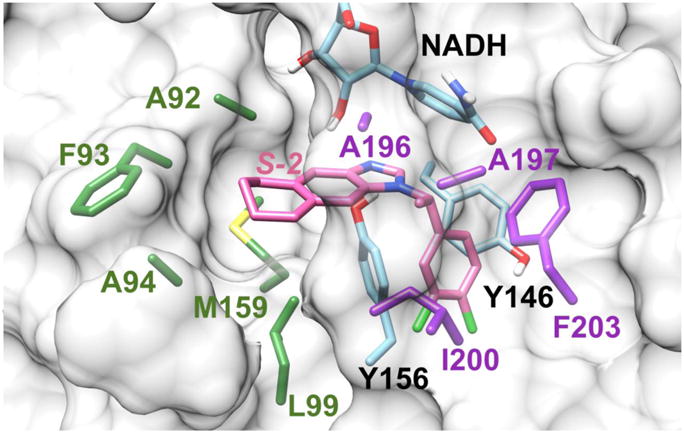
Binding mode of S-2 and FtFabI obtained by modifying the atoms of the ligand from the crystal structure of the FtFabI-benzimidazole complex (PDB IDs: 4J4T).19 The residues of A196, A197, I200, and F203 surrounding the chiral methyl are colored in purple. The residues (A92, F93, A94, L99, and M159) surrounding the six-member ring, which also build the larger tunnel-like pocket for benzimidazoles, are colored in green. Y146, Y156 and NADH are colored in cyan, and S-2 is shown in pink. The FtFabI enzyme is shown as a grey surface with only important residue sidechains shown, and with NADH partly shown to save space. The picture was prepared with Chimera1.10.2.42
