Abstract
The appearance of several disulfide bond isoforms in multiple cysteine containing venom peptides poses a significant challenge in their synthesis and purification under laboratory conditions. Recent experiments suggest that careful tuning of solvent and temperature conditions can propel the disulfide bond isoform equilibrium in favor of the most potent, native form. Certain aqueous ionic liquids (ILs) have proven significantly useful as solvents for this purpose, while exceptions have also been noted. To elucidate the molecular level origin behind such a preference, we report a detailed explicit solvent replica exchange molecular dynamics study of a conotoxin, AuIB, in pure water and four different aqueous IL solutions (~45–60% v/v). The ILs studied here are comprised of cations like 1-ethyl-3-methyl-imidazolium (Im21+) or 1-butyl-3-methyl-imidazolium (Im41+) coupled with either acetate (OAc−) or chloride (Cl−) as the counter anion. Our simulations unfold interesting features of the conformational spaces sampled by the peptide and its solvation in pure water and aqueous IL solutions. Detailed investigation into populations of the globular disulfide bond isoform of AuIB in aqueous IL solutions reveal distinct trends which might be related to the Hofmeister effect of the cation and anion of the IL and of specific interactions of the aqueous IL solutions with the peptide. In accordance with experimental observations, the aqueous [Im21][OAc] solution is found to promote the highest globular isoform population in AuIB.
Keywords: Ionic liquid, Conopeptide, REMD, Disulfide scrambling, Hofmeister effect
Introduction
Significant insight can be obtained into the vital biological networks present in higher organisms from the highly toxic venom molecules synthesized by them. Marine organisms are a source of innumerable venom peptides, a considerable number of which possess significant therapeutic promise. For example, conopeptides, the neurotoxic venoms produced by marine cone snails, are very aggressive and specific towards the human nervous system, attacking ion channels, receptors and transporters. Hence, they have been extensively researched for development as pain relievers or antinociceptive agents (Becker and Terlau 2008). Usually, conotoxins are ~10- to 40-residues long containing at least two or three disulfide bonds. The signature motifs of cysteine (Cys) and disulfide cross-links present in conopeptides help to maintain the proper three-dimensional structure essential for the toxin function and selectivity (Khoo et al. 2009, 2012; Góngora-Benítez et al. 2013). Additionally, the presence of multiple Cys residues pose the risk of formation of several disulfide bond isoforms when synthesized from linear precursors (Miloslavina et al. 2009; Heimer et al. 2014). The distinct isoforms possess unique structural and functional properties involving differences in potency and other attributes. However, the identification of proper disulfide pairing in a protein containing several Cys residues is exceedingly difficult, owing to incomplete disulfide bond formation and disulfide scrambling (Gupta et al. 2010). Scrambling of disulfide bond isoforms in such peptide venoms is a serious challenge towards their chemical synthesis and characterization, a technical hurdle often impossible to overcome in laboratories.
Conopeptide AuIB, a 15-residue toxin has been isolated from the marine cone snail species, Conus aulicus (Kaas et al. 2008). AuIB belongs to the ‘A’ superfamily of Conotoxins and is a specific non-competitive inhibitor of the α3–β4 subtype of the neuronal nicotinic acetylcholine receptors (nAChR) located in the mammalian postsynaptic membranes (Kaas et al. 2008, 2010, 2012; Grishin et al. 2010, 2013; Luo et al. 1998; Cho et al. 2000; Lebbe et al. 2014). It belongs to the framework-I and α4/6 subfamily of α– conotoxins, and the native globular isoform is noted for possessing a very robust α-helix stabilized by two disulfide bonds (Kaas et al. 2010). The α4/6 subfamily indicates the presence of 4 and 6 non-cysteine residues between the 2nd and 3rd and 3rd and 4th cysteine residues, respectively, in AuIB (Luo et al. 1998). The amino acid sequence of AuIB is given by GCCSYPPCFATNPDC. In native AuIB, the disulfide bridges connect the cysteine residues at positions 2–8 and 3–15, and the peptide has a globular structure. However, two other combinations of disulfide crosslinks are also possible, leading to the non-native ‘beads’ (2–3; 8–15) and ‘ribbon’ (2–15; 3–8) isoforms of AuIB. The numbers in parentheses denote the combination of the Cys residues in the peptide forming a S–S bond. AuIB is very special with respect to affinity and selectivity towards its receptor. Usually, conotoxins that target the α3–β4 nAChRs, also inhibit the α2–β3 subtype with comparable potency. However, AuIB selectively blocks only the α3–β4 subtype. AuIB is also unique because the non-native ribbon isoform is shown to be a more potent competitive inhibitor of the receptor than the native globular form (Lebbe et al. 2014).
It is well known that biomolecular structure and function are strongly influenced by the nature of the associated solvent and solvation dynamics (Bellissent-Funel et al. 2016; Ren et al. 2012). In particular, solvation and solvation dynamics of biomolecules in ionic liquids (ILs) are immensely diverse and have opened up a new era of experimental and computational studies, contributing greatly to the understanding of the molecular level details of the biomolecule-IL interface (Benedetto and Ballone 2016a, b; Schroder 2017). Ionic liquids have long been considered as environment conducive compared to conventional volatile organic compounds typically used as solvents for the synthesis of organic/bio-molecules, owing to their distinctive properties, such as very low vapor pressure, high thermal stability, non-flammability, non-combustibility and high ionic conductivity, to just name a few (Sheldon 2001; van Rantwijk and Sheldon 2007; Welton 1999). As solvents, ILs have been extensively studied for preserving biomolecules under extreme thermal conditions (Naushad et al. 2012). The technological efficacy of some biomolecules like proteins, nucleic acids and enzymes are also significantly enhanced in the presence of ILs as compared to conventional solvents (Sivapragasam et al. 2016). However, the dissolution ability of ILs is perhaps the most widely exploited property in the field of bio-molecular synthesis, extraction and preservation. ILs have been found to strongly affect protein structure and function by depleting the protein surface of water molecules, which are otherwise crucial in preserving the functional environment and polarity of the biomolecule. Especially, the IL anions having high H-bond basicity often significantly interfere with the native protein structure by tempering the existing H-bond networks (Yang 2012). With the help of a nucleophilic anion, an IL molecule can break through the strong H-bonding networks of hydrophobic biomolecules, subsequently dissolving them (Zhao et al. 2006; Zhao 2016). Recent studies have shown the remarkable ability of certain ILs to cleave disulfide bonds in proteins thereby inflicting significant changes in the protein secondary structure (Zhang et al. 2017; Guo et al. 2015). The ability of the ILs to dissolve a biomolecule further impacting the secondary structural elements of the protein depends strongly on the nature of the IL ions (Zhang et al. 2017). For the dissolution of hydrophobic peptides in particular, followed by their oxidative folding, the IL of choice is 1-ethyl-3-methyl-imidazolium acetate, [Im21][OAc]. Owing to its non-toxicity, low viscosity at room temperature and the presence of a strong nucleophilic and kosmotropic anion, OAc− together with a chaotropic cation Im21+, [Im21][OAc], is expected to stabilize the native folds of peptides/proteins (Miloslavina et al. 2009; Heimer et al. 2014; Zhao et al. 2006). However, recent findings indicate that all ILs essentially act as denaturants, and the size symmetric [Im21][OAc] is not an exception (Diddens et al. 2017). MD simulations probing preferential solvation of native and denatured states of a short hairpin peptide by Smiatek and co-workers have shown that OAc− strongly dehydrates peptides by accumulating in large numbers in the close vicinity of the biomolecule (Diddens et al. 2017; Lesch et al. 2015). In general, the presence of a kosmotropic anion and a chaotropic cation in [Im21][OAc] is believed to play an important role in protein stabilization (Zhao et al. 2006).
A groundbreaking work by Muttenthaler et al. showed that mutation of complementary cysteine pairs by selenocysteine pairs leads to the targeted native folding in at least five different classes of α-conotoxins (Muttenthaler et al. 2010). The enhanced hydrophobicity of the diselenide bond further increased the potency of these toxins towards their respective nicotinic acetylcholine receptors (Muttenthaler et al. 2010). In a more conventional chemical strategy, the complicated disulfide bond isoform population distribution can be significantly propelled towards a particular isoform by the careful choice of solvent and temperature conditions, as suggested by a recent series of experiments by Imhof and co-workers (Miloslavina et al. 2009; Heimer et al. 2014). The biocompatible ionic liquid [Im21][OAc] in particular helps the linear precursors of several conopeptides, like μ-SlllA, μ-PlllA and δ-EVlA, fold into the native isoform at room temperature in the absence of auxiliary redox agents and organic solvents (Heimer et al. 2014). Again, there are also reports in which [Im21][OAc] is shown to disfavor the formation of native folds in multiple cysteine-containing conopeptides, α-GI and CCAP-vil amongst others (Heimer et al. 2014; Lesch et al. 2015; Miloslavina et al. 2010). Thus, there appears to be a lacuna in understanding the particular reason why a given ionic liquid would favor a given disulfide bond isoform and the origin of the exceptions. This work is thus an attempt to recognize the reasons behind such preferences of ionic liquids in biasing the disulfide scrambling equilibrium towards a given isoform.
At this point, it is worth investigating if peptide sequence and disorder in secondary structures also play a decisive role in dictating whether an IL would favorably interact with it to promote native folding. A comprehensive review by Schröder gives a wonderful comparison of the various interactions that an IL may have with a peptide and the factors affecting the final fate of stabilization/destabilization (Schroder 2017). The stability and solubility of a peptide are strongly controlled by the nature of its interface with the solvent. Strong H-bonding between the IL ions and the peptide may result in large accumulations of ions in the vicinity of the peptide leading to its dehydration and subsequent destabilization (Schroder 2017; Lesch et al. 2015). An earlier impression was that the salts interfered with the structure of the water around the biomolecules, thereby affecting their solubility (Von Hippel and Wong 1964). However, subsequent studies have revealed that the ionic salts do not significantly alter the structure of bulk water. Rather, the solvated ions interact non-specifically with the biomolecular surface so as to alter the latter’s physico-chemical properties. Usually, the kosmotropic/chaotropic effect of the ions is only perceivable very close to the peptide, in its first solvation shell, and the impact fades away at distances from the peptide (Omta et al. 2003; Gurau et al. 2004; Zhang et al. 2005).
The dissolution of a biomolecule and the structure it exhibits in a complicated solvent system like that of aqueous ILs involve influences from multiple parameters like the polarity of the IL, the hydrophobicity of the IL ions, H-bonding donating and accepting abilities, etc. Apart from these, another well-explored property that helps in understanding the ionic hydration of biomolecules is the kosmotropicity or chaotropicity of the concerned ions, based on the classification made by Franz Hofmeister. Hofmeister, in his seminal work, showed that ions have differential abilities to “salt-in” or “salt-out” proteins from solutions and can be ordered into a series called the Hofmeister series (Gurau et al. 2004; Zhang et al. 2005; Hofmeister 1988; Tadeo et al. 2009). However, it is not always straightforward to correlate the salting-in/salting-out ions with chaotropes/kosmotropes, as pointed out by Zangi (2010). In fact, the salting-out phenomenon is an overall outcome of the tendency of proteins to correct folding and relative ordering, while enhanced solubilization with salting-in effect might lead to unfolding and subsequent aggregation or fibril formation (Benedetto and Ballone 2016). Ions that interact strongly and engage in H-bonding with water (like OAc−) can increase the water-structuring/order and are classified as ‘order makers’ or ‘kosmotropes’. Ions, which leave the water structure relatively unperturbed are called ‘chaotropes’. Cl− is marginally chaotropic (Zhao et al. 2006; Zhao 2006, 2016). Further, the combination of a kosmotropic anion (like OAc−) and a chaotropic cation (like Im21+) in an IL seems very significant as the former interacts strongly with water, expelling the latter from the strong anion–water network. The cation thus acts as a surfactant for the peptide molecule, and its concentration near the peptide surface may exceed that of the anion, regardless of the peptide charge, as confirmed by previous studies (Haberler et al. 2011, 2012; Klähn et al. 2011). Interestingly, the mean residence time of the anions near the peptide surface is usually much longer compared to the cations due to strong electrostatic and H-bonding interactions with specific positively charged amino acid residues in the peptide, especially arginine, lysine and histidine. The IL cations, on the other hand, only interact strongly with residues like glutamic acid and aspartic acid (Klähn et al. 2011). The destabilization of a peptide caused by an IL is most prominent in the solvent-exposed surface regions. The presence of coils makes the peptide more prone to attack by IL ions than helices and strands, which make the peptide structure compact and less solvent-exposed (Klähn et al. 2011). The sequence of the peptide and the disulfide bonds formed thereafter often dictate the solvent-exposed surface area of the peptide. Here probably lies the origin of the preference of an IL towards a given disulfide bond isoform in conopeptides, as the isoforms are often found to have different solvent-exposed surface areas (Dillon et al. 2008; Wan et al. 2011).
The present report addresses the phenomenon of disulfide scrambling in a neurotoxic conopeptide, AuIB, solvated in either water or four different aqueous IL solutions and the associated solvent-induced bias in the population distribution of different disulfide bond isoforms. The report also compares the general nature of preferential ionic solvation in AuIB and the effect of specific IL cations and anions on the peptide, as they are ranked in the Hofmeister series. Using explicit solvent replica exchange molecular dynamics (REMD) simulations, we investigate in detail the disulfide bond isoform equilibrium in AuIB responsible for the formation of a cocktail of isoforms in solution.
Simulation details
The starting coordinates of the ribbon isoform of AuIB (PDB ID: 1MXP) is obtained from the solution phase NMR structures of the peptide deposited in the RCSB Protein Data Bank (Berman et al. 2000; Dutton et al. 2002). All MD simulations employed NAMD as the molecular dynamics software (Kale et al. 1999; Phillips et al. 2005). The AMBER all atom force field developed by Cornel et al., and modified in CHARMM format, has been used to describe the bonding and non-bonding potentials of AuIB (Cornell et al. 1995). As mentioned in our previous works on AuIB (Roy and Lakshminarayanan 2016; Sajeevan 2017), this particular force field can parameterize the terminal as well as the non-terminal cysteine residues in AuIB, which may be either disulfide-bonded or reduced. The water molecules are explicitly modeled by the TIP3P force field as incorporated in CHARMM (Jorgensen et al. 1983; Jorgensen and Jenson 1998). The force field for the ionic liquid cation 1-alkyl-3-methylimidazolium is adapted from the work of Chaban et al. (2011), while the parameters for the anions OAc− and Cl− are taken from Charmm General Force Field, CGenFF (Vanommeslaeghe et al. 2009). The force fields used to model the peptide and the ionic liquids are compatible with each other.
We further modified the starting structure of the ribbon isoform of AuIB obtained from the Protein Data Bank by breaking the disulfide bond pairs, which is equivalent to the experimental condition of the complete reduction of the disulfide bonds in the peptide. This step is necessary as we aim to follow the structural evolution of a particular disulfide bond isoform into another, given that classical MD simulations cannot follow bond breaking and making processes. For the neat water simulations, the reduced peptide AuIB is solvated in a cubic box of TIP3P water containing 2850 water molecules and equilibrated for 40 ns in a NPT ensemble at 305 K and 1 atm. The final cubical box length is 44 Å along each axis. The center of the box is then moved to the center of mass of the peptide and all waters are wrapped. From this equilibrated box, a smaller cube of edge length 26 Å containing the peptide at the center is scooped out for further REMD simulations. The box prepared for REMD simulation contains the peptide along with the first hydration shell of 586 water molecules. For the simulations of the peptide in water-IL mixtures, 460 water and 52 IL molecules are equilibrated for 40 ns in a NPT ensemble at 305 K. The equilibrated cubical boxes have edge length ~30 Å. Thus, the aqueous IL solutions contained ~45–60% IL (v/v). The REMD simulations for peptide/water are run for 50 ns, while that for peptide/water-IL mixtures are run for 85 ns.
In all the simulations reported, a time step of 2 fs is used. The non-iterative SETTLE algorithm is used to keep waters and all bonds involving hydrogen rigid (Miyamoto and Kollman 1992). The REMD simulations for the peptide/water system consisting of 26 replicas have been run between temperatures 279–415 K with an exchange probability of ~0.5. For the peptide/water-IL system, 30 replicas have been simulated between 305 and 600 K with an exchange probability of ~0.3. The last 50 ns of peptide/aqueous IL systems and the last 40 ns for peptide/pure water system are considered for data analysis. Only the lowest temperature replica for each system (279 K for peptide/water and 305 K for peptide/water-IL) is considered for further structural analysis. Visualization of the structure and partial analysis of the replica exchange trajectories have been carried out using VMD (Humphrey et al. 1996). PLUMED (Bonomi et al. 2009; Tribello et al. 2014) has been used to calculate the S–S distances in AuIB. MATLAB (R2015b, v.8.0) has been used to analyze and plot the simulation data.
Results and discussions
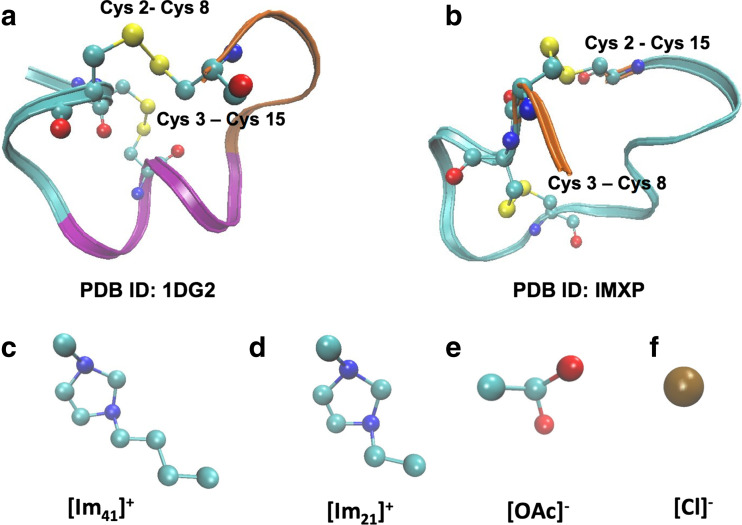
The structures of the globular and ribbon isoforms of AuIB along with the ionic liquid cations/anions studied in this work are depicted in Fig. 1. REMD simulations of peptides in explicit solvents are challenging owing to the strong masking effect of the solvent–solvent interaction potential on the total energy of the system during replica exchange (Periole and Mark 2007). When explicit solvent molecules are present, the peptide itself may not play the pivotal role in choosing its conformer during conformation sampling. Thus, it is essential to ensure that the peptide, the principal molecule of choice in the current study, is actually able to exploit its conformational energy to scan the coordinate space effectively during the REMD simulation. Fig. 2a, b shows the potential energy distributions of respectively, the total system (peptide plus solvent) and the only peptide for the AuIB/water-[Im21][OAc] case. While the distributions for the total energy are Gaussian as correctly expected from a temperature–REMD simulation, the ones with peptide alone are long-tailed. In this study, a high degree of overlap is observed between the peptide-only energy distributions, as observed previously by Periole and Mark (2007) However, the temperature effect is clearly seen in the widening of the distributions of the peptide potential energy (Fig. 2b) as the thermal energy of the replica is increased. Thus, it appears that, although the system is predominantly sensing solvent fluctuations during specific exchanges, it does seem to also percieve the conformations of the peptide when long trajectories are considered for analysis. Following the recommendations of Abraham and Gready (2008), we have assessed the mixing efficiency of our REMD simulations. In our case, for peptide/water and peptide/water-IL systems, respectively, the number of exchange attempts are 50,000 and 85,000, the swap interval is 1 ps and the total simulation lengths per replica are 50 ns and 85 ns. All these metrics should collectively lead to a reasonable REMD run.
Fig. 1.
Disulfide bond isoforms of AuIB. a Globular (Cys2–Cys8; Cys3–Cys15) and b ribbon isoform (Cys2–Cys15; Cys3–Cys8). Ionic liquid cations c 1-butyl-3-methylimidazolium, [Im41+] and d 1-ethyl-3-methylimidazolium, [Im21+] are depicted without hydrogens. The counter anions used are e acetate, [OAc−] and f chloride, [Cl−]
Fig. 2.
Internal energy distributions for AuIB/water-[Im21][OAc] system for 15 different replicas: a total system, b only peptide. The temperatures of the corresponding replicas are given
The principal aim behind this endeavor is to investigate how the different aqueous IL solutions help promote native globular folding in conotoxin AuIB. Towards this end, we have scanned the conformation space of AuIB in water and four aqueous IL solutions (Fig. 3). The purpose is to find the relative probability of obtaining three disulfide bond isoforms of AuIB and to understand the solvent influence on the process of disulfide scrambling. Since, in classical molecular dynamics force fields, bond making/breaking cannot be followed, the event of formation of a given disulfide bond is ascertained from the proximity of the sulfur (S) atoms of two cysteine residues under consideration, within a cutoff distance of 5 Å. Breaking the disulfide bonds through modeling of totally reduced AuIB is thus a necessary starting step in this study, as it attributes maximum freedom to the peptide to explore the conformation space and overcome the bias of the starting conformation. Figure 3 shows the S–S distance distributions for the three disulfide bond isoforms of AuIB in five solvation environments. The heat maps indicate the frame number density at a given S–S distance combination, unique for a given isoform. The plot uses 5000 snapshots belonging to the lowest temperature replica alone in each case, except for pure water, where 4000 snapshots are sampled. Essentially, the frames obtained within the 5 × 5 Å2 box are used for predicting the probability of observing the specific disulfide bond isoform in a given solvent. This includes those snapshots for which both the distance identifiers fall below a critical threshold value of 5 Å, which in turn increases the probability of the formation of a disulfide bond under oxidative conditions.
Fig. 3.
S–S distance distributions for the three disulfide bond isoforms of AuIB (B beads, G globular, R ribbon) in neat water and four aqueous IL solutions. The plots show the distribution of snapshots sampled in the lowest temperature replica alone and contain 5000 scattered points for the aqueous IL solutions and 4000 points for pure water. The heat maps show the density of snapshots for a given combination of the distance identifiers, dij and dkl
Figure 3 reveals two features of the frequency of occurrence for the three disulfide bond isoforms. Firstly, it is worth knowing how many snapshots actually fall within the 5 × 5 Å2 region for a given isoform in a given solvent system. This number might be reminiscent of the folding propensity of AuIB in that particular solvation environment. Secondly, the normalized relative probability of obtaining three isoforms in a given solvent system is important to understand the bias that the solvation environment imputes towards a particular isoform. Table 1 lists the actual number of snapshots considered for frequency calculation in all the systems. From the data, it is evident that, while the overall folding yield for all the three disulfide bond isoforms, estimated by the snapshot hits within 5 × 5 Å2 of their corresponding distance identifiers, is highest for AuIB solvated in aqueous [Im21][Cl] solution, the solvent bias towards the native globular isoform is highest in the aqueous [Im21][OAc] solution. The observation that the aqueous [Im21][OAc] solution supports native folds in several conopeptides is supported by experimental results (Miloslavina et al. 2009; Heimer et al. 2014; Tietze et al. 2012). It is also worth noting that, despite starting the simulations from the modified crystal structure of the ribbon isoform of AuIB (PDB: 1MXP), at the end of the REMD simulation, the bias of the starting structure is not at all noticeable. Significant population redistribution from the ribbon isoform of AuIB to the globular or beads isoforms is noted in all the five solvent systems. The relative proportions of the different isoforms of AuIB in water and the four aqueous IL solutions are tabulated in Table 1. Our simulations indicate that ionic liquid solutions containing Cl− as the anion barely show a bias towards a particular isoform of AuIB. All the three isoform populations are seen appreciably populating the aqueous [Im21][Cl] and [Im41][Cl] solutions. These mixtures are also efficient in promoting the overall folding yield of the totally reduced peptide (as estimated by the total number of snapshots coming within 5 × 5 Å2 of all the isoforms). On the other hand, solutions containing OAc− as the anion show a specific bias towards the native globular form. However, the overall folding propensity in these solutions appears to be low.
Table 1.
Normalized probabilities and yields of formation of a disulfide bond isoform in AuIB solvated in either water or aqueous IL solutions
| Solvent | IL viscosity B-coefficient (dm3 mol−1) |
Globular (%) | Ribbon (%) | Beads (%) |
|---|---|---|---|---|
| Water | – | 60.9 (53) | 31.3 (28) | 7.8 (7) |
| Water-[Im21][OAc] | −0.338 | 83.3 (50) | 0 (0) | 16.6 (10) |
| Water-[Im41][OAc] | −0.457 | 64.8 (35) | 1.9 (1) | 33.3 (18) |
| Water-[Im21][Cl] | −0.485 | 46.5 (66) | 21.8 (31) | 31.7 (45) |
| Water-[Im41][Cl] | −0.604 | 36.3 (28) | 16.9 (13) | 46.8 (36) |
For peptide/water simulations, 279 K is the lowest temperature replica. and for peptide/aqueous IL system it is 305 K. Data from only the lowest temperature replica of each system are used for the analysis. The numbers in parentheses are the actual number of snapshots from the lowest temperature replica coming within the cutoff of 5 Å for both the distance identifiers unique for a given isoform
To quantify the extent of overall kosmotropicity of an IL, we have used the thermodynamic viscosity B-coefficients (Fig. 4; Table 1). The viscosity B-coefficients are obtained from the Jones–Dole empirical equation of the relative viscosities of electrolyte solutions as a function of their concentrations (Jenkins and Marcus 1995). Work by Zhao and co-workers shows correlations of the viscosity B-coefficient of an ion to its hydration radius (Zhao et al. 2006; Zhao 2006, 2016). In Fig. 4, we have tried to find a relationship between the effective viscosity B-coefficient of the IL (given by the difference in anion and cation viscosity B-coefficients) and the percentage of globular or beads isoforms of AuIB in the different aqueous IL solutions.The plot shows that aqueous IL solutions with a higher effective B-coefficient value favor more globular and fewer bead isoforms for AuIB. Additional simulations with different ion combinations are required to establish this observation.
Fig. 4.
Correlation between the percentage of disulfide bond isoforms (black square globular; red triangle beads) and the difference in the Jones–Dole viscosity B-coefficients of anion and cation (in dm3 mol−1). The viscosity B-coefficient values for the cations and anions are taken from Zhao (2006), following equations 4 and 5
In order to have a better understanding of the systems, we have studied the structure of the solvation shell of AuIB in the five systems. Figure 5 shows the radial distribution functions, g(r), of the aqueous IL components around the peptide and those between water and the IL ions in the five solvation environments. All distributions, except that for OW–OW (OW: oxygen atom of water) are calculated within the first solvation shell of 7 Å around AuIB considering only the lowest temperature replica in each case. The peptide–cation g(r) plots (Fig. 5a) show that the distribution of IL cations around AuIB are almost the same for Im21+ and Im41+, with the latter showing slightly higher occupancy around the peptide, irrespective of its counterion, OAc− or Cl−. However, the peptide–anion g(r) plots (Fig. 5b) show significant variations with respect to shape. It appears that the aqueous IL solutions containing OAc− as the anion show noticeably higher anion accumulation around the peptide than those containing Cl− as the counterion. In fact, the Cl− ions seem to be selectively excluded from being very close to the peptide. This observation is reminiscent of the simulation results of Smiatek and co-workers, as mentioned previously (Diddens et al. 2017; Lesch et al. 2015). The exceptional affinity of OAc− ion for AuIB dominates the first solvation shell of the peptide. This interaction is attributed to substantial H-bonding between the amino acid residues of AuIB and OAc−, as discussed later.
Fig. 5.
Radial distribution functions involving AuIB and the solvating species in neat water and four aqueous IL solutions: a peptide–cation; b peptide–anion; c water–cation; d water–anion; (e) between oxygen atoms of water within 2.5 Å from the peptide and f peptide–water. All the distributions except (e) are calculated within 7 Å from the peptide surface
Since aqueous ILs solvating a biomolecule always invoke the manifestation of the Hofmeister effect (Hofmeister 1988), we would like to explore how the different combinations of cation and anion in the aqueous IL perturb the water structures in the vicinity of AuIB. The g(r) plots between water and IL ions in the close vicinity of the peptide (Fig. 5c, d) indicate the subtle effects of distress in the water structure caused by an ionic species. The effect fades away as the distribution encompasses larger radius around the peptide (Omta et al. 2003). The plots indicate that the cations minimally interact with water in the vicinity of the peptide. However, if the cations are associated with OAc− as the counterion, they interact relatively more with water than in presence of Cl− as the counterion. The water–anion g(r) plots (Fig. 5d), on the other hand, are very contrasting for OAc− and Cl−. The OAc− ion involves in significantly strong specific interactions with water having a notably close penetration distance. The Cl− ions, on the other hand, have a single site and are thus unable to come very close to the water molecules. This is also evident to some extent from the OW–OW g(r) plots (Fig. 5e). The selected water molecules lie within a distance of 2.5 Å from the peptide and the g(r) plots are normalized with respect to the bulk. These distributions are overall pretty close to each other, within 3 Å, beyond which a very subtle decrease in the height of the distributions is noticed for the ILs. The chloride-containing ILs, on the other hand, show similar distributions as neat water. This probably indicates that the OAc− ions interfere with the strong H-bonding network of water clusters in the vicinity of the peptide by acting as a strong kosmotrope, while Cl− ions, being rather chaotropic, retain the water network relatively unperturbed.
The last g(r) plot of this series (Fig. 5f) depicts how the water molecules are organized within 7 Å of AuIB. This figure shows a clear dehydration effect in the vicinity of the peptide induced by the water-IL mixtures containing OAc− as the anion. The Cl−-containing aqueous ILs, on the other hand, helped retain the water layer on the peptide surface to an extent even higher than that observed in case of neat water, possibly due to strong electrostatic interaction of the IL ions with the peptide and also their close association with water.
When we discuss specific interactions of the solvent components with AuIB, the analysis of intra-peptide and peptide-solvent H-bonds comes inadvertently. Figure 6 shows H-bonding propensities between different combinations of peptide and solvent components. From Fig. 6a, it appears that the Cl−-containing ionic liquids promote more intra-peptide H-bonds than the OAc−-containing ones. The aqueous ILs containing OAc− ions engage in strong peptide–anion H-bonding, as evident from the distributions in Fig. 6c, thereby breaking the intra-peptide H-bonds. The entire H-bonding effect seems to be dominated by the IL anions in these four water-IL mixtures while the cationic counterparts play a small role. Overall, the number of intermolecular peptide–solvent H-bonds is higher in the OAc− ion-containing solvent mixtures than that in the chloride-containing ones. However, these are far less than that found in neat water. Fig. 6d shows the peptide–water H-bonds for all the systems involving only polar atoms (N, O, S) in AuIB. Notably, AuIB dissolved in aqueous IL solutions containing OAc− forms a higher number of H-bonds with the solvent than that found in the Cl−-containing water-IL mixtures. This is attributed to much higher involvement of the OAc− ions in H-bonding with both water and peptide. However, these numbers are significantly less when compared to the peptide–water H-bonding in neat water.
Fig. 6.
Intra-peptide and peptide-solvent H-bond distributions: a intra-peptide; b peptide–all components of the solvent/solvent mixture; c peptide–anion and d peptide–water. Except for (d), for all the other plots, the calculations are carried out by considering that a hydrogen bond is formed between any atom with a hydrogen bonded to it (the donor, D) and another atom (the acceptor, A) provided that the distance D–A is less than the cutoff distance of 4 Å and the angle D–H–A is less than the cutoff angle of 40°. For (d) only the polar atoms (N, O, S, F) are specifically considered in the H-bond calculation
The effect of the aqueous IL solutions on peptide compression can be ascertained by analyzing the backbone length of the peptide from the lowest temperature replica of AuIB in all five solvent systems (Fig. 7). This plot shows an interesting bimodal behavior with a cross-over at ~17.5 Å. The aqueous IL solutions containing Cl− as the anion favor smaller backbone lengths as compared to OAc−. Further, the mixtures containing Im21+ as the cation consistently favor smaller lengths as compared to Im41+. The aqueous [Im21][Cl] solution shows an exceptionally high preference for the compact peptide with an average backbone length of ~15–16 Å. There is also a minor peak at ~ 20–22 Å, corresponding to the relatively open conformer of the peptide. The fundamental cause of the difference in backbone lengths in the two populations are the closing-in/opening-up motions of the N- and C-terminal domains of the peptide, which either move closer or fall apart during the course of the conformation search. In Fig. 7b, we compare the same distributions for all the snapshots which satisfy the criteria of two S–S bond formation and are within the 5 × 5 Å2 distance cutoffs. The last plot (Fig. 7c) shows the area normalized version of Fig. 7b, which avoids the effect of different number of snapshots falling within the cutoff in different mixtures. From Fig. 7b, c, the preferences for backbone length and the overall folding propensity of AuIB are clearly observed for the two sets of OAc−- and Cl−- containing aqueous IL solutions.
Fig. 7.
Distributions of the peptide backbone length of AuIB in neat water and four aqueous IL solutions for: a all snapshots belonging to the lowest temperature replica; b snapshots belonging to the lowest temperature replica and falling within the 5 × 5 Å2 cutoff for the distance identifiers of either globular, beads or ribbon isoforms of AuIB; c the same as in (b), but with the area normalized
Lastly, we need to compare the solvation energies of the peptide in these four water-IL mixtures. Figure 8 shows the weightage of electrostatic and van der Waals components to the total solvation energy. The analysis shows the striking lack of stabilization of the peptide AuIB in the aqueous [Im21][Cl] solution in comparison to the other solutions under consideration (Table 2). A close analysis reveals that, for the OAc−-containing aqueous IL solutions, the weightage of anion electrostatics is more than that of the cation, while it is reversed for the Cl−-containing ILs. The vdW contributions to the total nonbonding energy are more for the cations than the anions for their evidently larger size as compared to the single site Cl− and slightly bigger OAc− ions. The exceptionally low contribution to the overall non-bonding energy from water in aqueous [Im21][Cl] solution is probably the reason why the peptide is so compact in this system. However, this effect is not visible for the aqueous [Im41][Cl] solution and seems unique for the combination of Im21+ cation and Cl− anion. Smiatek and co-workers recently observed this effect for the aqueous [Im21][Cl] solution on the native and denatured states of a short hairpin peptide (Diddens et al. 2017). They further observed that size-symmetric ILs like [Im21][OAc] tend to be stronger denaturants than size-asymmetric ILs and exhibit marked accumulation of ions around the peptide, further supporting our observations for AuIB. The ion accumulation observed in the case of aqueous ILs containing the OAc− ion induced a strong dehydration effect on the peptide, thereby favoring unfolded structures (Diddens et al. 2017; Canchi and Garcia 2013). Additionally, small non-coordinating ions like Cl− and BF4− are found to be preferentially excluded from close distances around peptide conformations. All these observations are qualitatively in line with our simulation results.
Fig. 8.
Non-bonding potential energies (van der Waals and electrostatic) of interaction between AuIB and the components of the solvent mixture: water (W), anion (A), cation (C) and total solvent (T)
Table 2.
Non-bonding interaction energies (kcal mol−1) between AuIB and the different solvent components (IL ions and water) in water and four aqueous IL solutions
| Solvent | Peptide–cation | Peptide–anion | Peptide–water | Peptide–solvent | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Eelec | EvdW | Etot | Eelec | EvdW | Etot | Eelec | EvdW | Etot | Eelec | EvdW | Etot | |
| Water | – | – | – | – | – | – | −790 | −61 | −851 | −790 | −61 | −851 |
| Water-[Im21][OAc] | −216 | −59 | −275 | −292 | −15 | −307 | −247 | −17 | −265 | −756 | −92 | −848 |
| Water-[Im41][OAc] | −208 | −71 | −279 | −325 | −12 | −338 | −235 | −12 | −248 | −769 | −96 | −865 |
| Water-[Im21][Cl] | −184 | −66 | −251 | −98 | −3 | −101 | −47 | 18 | −28 | −330 | −50 | −380 |
| Water-[Im41][Cl] | −192 | −80 | −273 | −166 | −2 | −168 | −310 | −13 | −324 | −670 | −96 | −766 |
Conclusions and future works
Molecular dynamics simulation using non-reactive and non-polarizable force fields does not give the complete picture of the disulfide scrambling process, as it cannot realistically model the bond-breaking and -making processes. To overcome this problem, in the current study, we have employed the S–S cutoff distance criteria to ascertain whether a given disulfide bond is formed. This information is further used to calculate the frequency of the isoforms observed across the REMD trajectories. Advanced reactive force fields like ReaxFF would be more pertinent to study such phenomena related to disulfide bonds (Monti et al. 2013; Müller and Hartke 2016). Additional studies are also required to find a direct correlation of the disulfide bond isoform frequencies with their structural attributes in the five-solvent systems. The authors believe that the solvent-exposed surface area of the respective isoforms plays a decisive role in dictating the preference of the solvent. In a separate experiment involving the disulfide bond isoforms of IgG2 monoclonal antibodies, it was shown that the different isoforms possess diverse solvent-exposed surface areas and also differ in terms of hydrophobicity (Dillon et al. 2008). This variation in their exposed surface areas indeed plays a crucial role in dictating their receptor binding preference. In fact, the IgG2 isoforms were shown to scramble in a “blood-like” environment and the distribution of isoforms is believed to affect the in vivo activity of human IgG2 (Dillon et al. 2008). Here lies the significance of understanding the disulfide scrambling process and the associated equilibrium in such multiple cysteine-containing peptides/proteins. This work is an initial attempt to understand the effect of ionic hydration on the disulfide bond isoform equilibrium very often found in venom toxins. By combining two cations of the imidazolium type, Im21+ (chaotropic) and Im41+ (kosmotropic), with two anions, OAc− (kosmotropic) and Cl− (chaotropic), four different aqueous IL solutions are created to understand how specific and general ionic solvation help drive the population of the native globular isoform in AuIB. Our simulations indicate that, although peptide solvation is predominantly influenced by the anions, the overall kosmotropicity of the IL is a decisive factor in regulating the disulfide bond isoform equilibrium.
This article primarily deals with the structural aspect of the scrambling phenomenon. However, the kinetic feature of the problem, answering questions like the rates of inter-conversion between the different local and global conformations of AuIB and their kinetic coupling could also be very important. REMD simulations in conjunction with methods developed by Buchete and Hummer involving a coarse master equation approach could possibly handle the associated dynamics happening over different time scales (Buchete and Hummer 2008). Additional works exploiting all the aforementioned ideas are underway with diverse IL and peptide combinations and with reactive force fields to gain a general understanding of the mechanism by which ILs influence the disulfide scrambling process.
Acknowledgements
DR sincerely acknowledges the encouragement and support received from Professor Mark Maroncelli of the Pennsylvania State University. The authors are thankful to Dr. Debashis Barik, University of Hyderabad, for his kind help in providing computation time. KAS and DR are grateful to the Department of Science & Technology, India (SERB, YSS/2014/000301), for funding. Authors also gratefully acknowledge support from Department of Science and Technology, India, for the FIST grant SR/FST/CSI-240/2012.
Compliance with ethical standards
Conflict of interest
Karuna Anna Sajeevan declares that she has no conflict of interest. Durba Roy declares that she has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Footnotes
This article is part of a Special Issue on ‘Ionic Liquids and Biomolecules’ edited by Antonio Benedetto and Hans-Joachim Galla.
References
- Abraham MJ, Gready JE (2008) Ensuring mixing efficiency of replica-exchange molecular dynamics simulations. J Chem Theory Comput 4(7):1119–1128 [DOI] [PubMed]
- Becker S, Terlau H (2008) Toxins from cone snails: properties, applications and biotechnological production. Appl Microbiol Biotechnol 79:1–9 [DOI] [PMC free article] [PubMed]
- Bellissent-Funel M-C, Hassanali A, Havenith M, Henchman R, Pohl P, Sterpone F, van der Spoel D, Xu Y, Garcia AE (2016) Water determines the structure and dynamics of proteins. Chem Rev 116(13):7673–7697 [DOI] [PMC free article] [PubMed]
- Benedetto A, Ballone P (2016a) Room temperature ionic liquids meet biomolecules: a microscopic view of structure and dynamics. ACS Sustain Chem Eng 4(2):392–412
- Benedetto A, Ballone P (2016b) Room temperature ionic liquids interacting with bio-molecules: an overview of experimental and computational studies. Philos Mag 96(7–9):870–894
- Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242 [DOI] [PMC free article] [PubMed]
- Bonomi M, Branduardi D, Bussi G, Camilloni C, Provasi D, Raiteri P, Donadio D, Marinelli F, Pietrucci F, Broglia RA, Parrinello M (2009) PLUMED: A portable plugin for free-energy calculations with molecular dynamics. Comput Phys Commun 180(10):1961–1972
- Buchete NV, Hummer G (2008) Coarse master equations for peptide folding dynamics. J Phys Chem B 112(19):6057–6069 [DOI] [PubMed]
- Canchi DR, Garcia AE (2013) Cosolvent effects on protein stability. Annu Rev Phys Chem 64:273–293 [DOI] [PubMed]
- Chaban VV, Voroshylovab IV, Kaluginb ON (2011) A new force field model for the simulation of transport properties of imidazolium-based ionic liquids. Phys Chem Chem Phys 13:7910–7920 [DOI] [PubMed]
- Cho J-H, Mok KH, Olivera BM, McIntosh JM, Park K-H, Han K-H (2000) Nuclear magnetic resonance solution conformation of alpha-conotoxin AuIB, an alpha(3)beta(4) subtype-selective neuronal nicotinic acetylcholine receptor antagonist. J Biol Chem 275(12):8680–8685 [DOI] [PubMed]
- Cornell WD, Cieplak P, Bayly CI, Gould IR, JKM M, Ferguson DM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J Am Chem Soc 117:5179–5197
- Diddens D, Lesch V, Heuerab A, Smiatek J (2017) Aqueous ionic liquids and their influence on peptide conformations: denaturation and dehydration mechanisms. Phys Chem Chem Phys 19:20430–20440 [DOI] [PubMed]
- Dillon TM, Ricci MS, Vezina C, Flynn GC, Liu YD, Rehder DS, Plant M, Henkle B, Li Y, Deechongkit S, Varnum B, Wypych J, Balland A, Bondarenko PV (2008) Structural and functional characterization of disulfide isoforms of the human IgG2 subclass. J Biol Chem 283:16206–16215 [DOI] [PMC free article] [PubMed]
- Dutton JL, Bansal PS, Hogg RC, Adams DJ, Alewood PF, Craik DJ (2002) A new level of conotoxin diversity, a non-native disulfide bond connectivity in alpha-Conotoxin AuIB reduces structural definition but increases biological activity. J Biol Chem 277:48849–48857 [DOI] [PubMed]
- Góngora-Benítez M, Tulla-Puche J, Albericio F (2013) Constella™(EU)-Linzess™(USA): the last milestone in the long journey of the peptide linaclotide and its implications for the future of peptide drugs. Future Med Chem 5(3):291–300 [DOI] [PubMed]
- Grishin AA, Wang CI, Muttenthaler M, Alewood PF, Lewis RJ, Adams DJ (2010) Alpha-conotoxin AuIB isomers exhibit distinct inhibitory mechanisms and differential sensitivity to stoichiometry of alpha3beta4 nicotinic acetylcholine receptors. J Biol Chem 285(29):22254–22263 [DOI] [PMC free article] [PubMed]
- Grishin AA, Cuny H, Hung A, Clark RJ, Brust A, Akondi K, Alewood PF, Craik DJ, Adams DJ (2013) Identifying key amino acid residues that affect alpha-conotoxin AuIB inhibition of alpha3-beta4 nicotinic acetylcholine receptors. J Biol Chem 288(48):34428–34442 [DOI] [PMC free article] [PubMed]
- Guo M, Zhai Y, Guo C, Liu Y, Tang D, Pan Y (2015) A new strategy to determine the protein mutation site using matrix-assisted laser desorption ionization in-source decay: Derivatization by ionic liquid. Anal Chim Acta 865:31–38 [DOI] [PubMed]
- Gupta K, Kumar M, Balaram P (2010) Disulfide bond assignments by mass spectrometry of native natural peptides: cysteine pairing in disulfide bonded conotoxins. Anal Chem 82:8313–8319 [DOI] [PubMed]
- Gurau MC, Lim S-M, Castellana ET, Albertorio F, Kataoka S, Cremer PS (2004) On the mechanism of the Hofmeister effect. J Am Chem Soc 126:10522–10523 [DOI] [PubMed]
- Haberler M, Schroder C, Steinhauser O (2011) Solvation studies of a zinc finger protein in hydrated ionic liquids. Phys Chem Chem Phys 13(15):6955–6969 [DOI] [PMC free article] [PubMed]
- Haberler M, Schröder C, Steinhauser O (2012) Hydrated ionic liquids with and without solute: the influence of water content and protein solutes. J Chem Theory Comput 8(10):3911–3928 [DOI] [PubMed]
- Heimer P, Tietze AA, Bohm M, Giernoth R, Kuchenbuch A, Stark A, Leipold E, Heinemann SH, Kandt C, Imhof D (2014) Application of room-temperature aprotic and protic ionic liquids for oxidative folding of cysteine-rich peptides. Chembiochem 15(18):2754–2765 [DOI] [PubMed]
- Hofmeister F (1988) Zur lehre von der wirkung der salze. Arch Exp Pathol Pharmakol 24:247–260
- Humphrey W, Dalke A, Schulten K (1996) VMD - visual molecular dynamics. J Molec Graph 14:33–38 [DOI] [PubMed]
- Jenkins HDB, Marcus Y (1995) Viscosity BCoefficients of ions in solution. Chem Rev 95:2695–2724
- Jorgensen WL, Jenson C (1998) Temperature dependence of TIP3P, SPC, and TIP4P water from NPT monte carlo simulations: seeking temperatures of maximum density. J Comput Chem 19(10):1179–1186
- Jorgensen WL, Chandrasekhar J, Madura JD (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
- Kaas Q, Westermann JC, Halai R, Wang CK, Craik DJ (2008) ConoServer, a database for conopeptide sequences and structures. Bioinformatics 24(3):445–446 [DOI] [PubMed]
- Kaas Q, Westermann J-C, Craik DJ (2010) Conopeptide characterization and classifications: an analysis using ConoServer. Toxicon 55(8):1491–1509 [DOI] [PubMed]
- Kaas Q, Yu R, Jin AH, Dutertre S, Craik DJ (2012) ConoServer: updated content, knowledge, and discovery tools in the conopeptide database. Nucleic Acids Res 40(Database Issue):D325–D330 [DOI] [PMC free article] [PubMed]
- Kale L, Skeel R, Bhandarkar M, Brunner R, Gursoy A, Krawetz N, Phillips J, Shinozaki A, Varadarajan K, Schulten K (1999) NAMD2: Greater scalability for parallel molecular dynamics. J Comput Phys 151:283–312
- Khoo KK, Feng Z-P, Smith BJ, Zhang M-M, Yoshikami D, Olivera BM, Bulaj G, Norton RS (2009) Structure of the analgesic μ-Conotoxin KIIIA and effects on the structure and function of disulfide deletion. Biochemistry 48:1210 [DOI] [PMC free article] [PubMed]
- Khoo KK, Gupta K, Green BR, Zhang M-M, Watkins M, Olivera BM, Balaram P, Yoshikami D, Bulaj G, Norton RS (2012) Distinct disulfide isomers of μ-Conotoxins KIIIA and KIIIB block voltage-gated sodium channels. Biochemistry 51(49):9826–9835 [DOI] [PMC free article] [PubMed]
- Klähn M, Lim GS, Seduraman A, Wu P (2011) On the different roles of anions and cations in the solvation of enzymes in ionic liquids. Phys Chem Chem Phys 13(4):1649–1662 [DOI] [PubMed]
- Lebbe EKM, Peigneur S, Wijesekara I, Tytgat J (2014) Conotoxins targeting nicotinic acetylcholine receptors: an overview. Marine Drugs 12:2970–3004 [DOI] [PMC free article] [PubMed]
- Lesch V, Heuer A, Tatsis VA, Holm C, Smiatek J (2015) Peptides in the presence of aqueous ionic liquids: tunable co-solutes as denaturants or protectants? Phys Chem Chem Phys 17:26049–26053 [DOI] [PubMed]
- Luo S, Kulak JM, Cartier GE, Jacobsen RB, Yoshikami D, Olivera BM, McIntosh JM (1998) Alpha-conotoxin AuIB selectively blocks alpha3 beta4 nicotinic acetylcholine receptors and nicotine-evoked norepinephrine release. J Neurosci 18:8571–8579 [DOI] [PMC free article] [PubMed]
- Miloslavina AA, Leipold E, Kijas M, Stark A, Heinemann SH, Imhof D (2009) A room temperature ionic liquid as convenient solvent for the oxidative folding of conopeptides. J Peptide Sci 15(2):72–77 [DOI] [PubMed]
- Miloslavina A, Ebertb C, Tietze D, Ohlenschläger O, Englert C, Görlach M, Imhof D (2010) An unusual peptide from Conus villepinii: synthesis, solution structure, and cardioactivity. Peptides 31:1292–1300 [DOI] [PubMed]
- Miyamoto S, Kollman PA (1992) SETTLE: an analytical version of the SHAKE and RATTLE algorithm for rigid water models. J Comput Chem 13(8):952–962
- Monti S, Corozzi A, Fristrup P, Joshi KL, Shin YK, Oelschlaeger PC, van Duin AC, Barone V (2013) Exploring the conformational and reactive dynamics of biomolecules in solution using an extended version of the glycine reactive force field. Phys Chem Chem Phys 15(36):15062–15077 [DOI] [PubMed]
- Müller J, Hartke B (2016) ReaxFF reactive force field for disulfide mechanochemistry, fitted to multireference ab initio data. J Chem Theory Comput 12(8):3913–3925 [DOI] [PubMed]
- Muttenthaler M, Nevin ST, Grishin AA, Ngo ST, Choy PT, Daly NL, Hu S-H, Armishaw CJ, Wang C-IA, Lewis RJ, Martin JL, Noakes PG, Craik DJ, Adams DJ, Alewood PF (2010) Solving the a-Conotoxin folding problem: efficient selenium-directed on-resin generation of more potent and stable nicotinic acetylcholine receptor antagonists. J Am Chem Soc 132:3514–3522 [DOI] [PubMed]
- Naushad M, Alothman ZA, Khan AB, Ali M (2012) Effect of ionic liquid on activity, stability, and structure of enzymes: a review. Int J Biol Macromol 51(4):555–560 [DOI] [PubMed]
- Omta AW, Kropman MF, Woutersen S, Bakker HJ (2003) Negligible effect of ions on the hydrogen-bond structure in liquid water. Science 301:347–349 [DOI] [PubMed]
- Periole X, Mark AE (2007) Convergence and sampling efficiency in replica exchange simulations of peptide folding in explicit solvent. J Chem Phys 126:014903 [DOI] [PubMed]
- Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kale L, Schulten K (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26:1781–1802 [DOI] [PMC free article] [PubMed]
- Ren P, Chun J, Thomas DG, Schnieders MJ, Marucho M, Zhang J, Baker NA (2012) Biomolecular electrostatics and solvation: a computational perspective. Q Rev Biophys 45(4):427–491 [DOI] [PMC free article] [PubMed]
- Roy D, Lakshminarayanan M (2016) Scrambling of disulfide bond scaffolds in neurotoxin AuIB: a molecular dynamics simulation study. Biopolymers (Pept Sc) 106(2):196–209 [DOI] [PubMed]
- Sajeevan KA (2017) Roy, D. Temperature dependent molecular dynamics study reveals an ionic liquid induced 310- to α-helical switch in a neurotoxin. Pept Sci 108(3):e23009 [DOI] [PubMed]
- Schroder C (2017) Proteins in ionic liquids: current status of experiments and simulations. Top Curr Chem 375:25 [DOI] [PMC free article] [PubMed]
- Sheldon R (2001) Catalytic reactions in ionic liquids. Chem Commun 0(23):2399–2407 [DOI] [PubMed]
- Sivapragasam M, Moniruzzaman M, Goto M (2016) Recent advances in exploiting ionic liquids for biomolecules: solubility, stability and applications. Biotechnol J 11(8):1000–1013 [DOI] [PubMed]
- Tadeo X, Lopez-Mendez B, Castano D, Trigueros T, Millet O (2009) Protein stabilization and the hofmeister effect: the role of hydrophobic solvation. Biophys J 97:2595–2603 [DOI] [PMC free article] [PubMed]
- Tietze AA, Heimer P, Stark A, Imhof D (2012) Ionic liquid applications in peptide chemistry: synthesis, purification and analytical characterization processes. Molecules 17(4):4158–4185 [DOI] [PMC free article] [PubMed]
- Tribello GA, Bonomi M, Branduardi D, Camilloni C, Bussi G (2014) Plumed 2: new feathers for an old bird. Comput Phys Commun 185(2):604–613
- van Rantwijk F, Sheldon RA (2007) Biocatalysis in ionic liquids. Chem Rev 107(6):2757–2785 [DOI] [PubMed]
- Vanommeslaeghe K, Hatcher E, Acharya C, Kundu S, Zhong S, Shim J, Darian E, Guvench O, Lopes P, Vorobyov I, Mackerell AD Jr (2009) CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J Comput Chem 31(4):671–690 [DOI] [PMC free article] [PubMed]
- Von Hippel PH, Wong KY (1964) Neutral salts: the generality of their effects on the stability of macromolecular conformations. Science 145:577–580 [DOI] [PubMed]
- Wan X, Kumar S, Singh SK (2011) Disulfide scrambling in IgG2 monoclonal antibodies: insights from molecular dynamics simulations. Pharm Res 28(12):3128–3144 [DOI] [PubMed]
- Welton T (1999) Room-temperature ionic liquids. Solvents for synthesis and catalysis. Chem Rev 99:2071–2083 [DOI] [PubMed]
- Yang Z (2012) Ionic liquids and proteins: academic and some practical interactions. In: de María PD (ed) Ionic liquids in biotransformations and organocatalysis: solvents and beyond. Wiley, New York, pp 15–71
- Zangi R (2010) Can salting-in/salting-out ions be classified as chaotropes/kosmotropes? J Phys Chem B 114(1):643–650 [DOI] [PubMed]
- Zhang Y, Furyk S, Bergbreiter DE, Cremer PS (2005) Specific ion effects on the water solubility of macromolecules: PNIPAM and the hofmeister series. J Am Chem Soc 127:14505–14510 [DOI] [PubMed]
- Zhang Z, Nie Y, Zhang Q, Liu X, Tu W, Zhang X, Zhang S-J (2017) Quantitative change of disulfide bond and microstructure variation of the regenerated wool keratin from various ionic liquids. ACS Sustain Chem Eng 5(3):2614–2622
- Zhao H (2006) Are ionic liquids kosmotropic or chaotropic? An evaluation of available thermodynamic parameters for quantifying the ion kosmotropicity of ionic liquids. J Chem Technol Biotechnol 81:877–891
- Zhao H (2016) Protein stabilization and enzyme activation in ionic liquids: specific ion effects. J Chem Technol Biotechnol 91:25–50 [DOI] [PMC free article] [PubMed]
- Zhao H, Olubajo O, Song Z, Sims AL, Person TE, Lawal RA, Holley LA (2006) Effect of kosmotropicity of ionic liquids on the enzyme stability in aqueous solutions. Bioorg Chem 34:15–25 [DOI] [PubMed]