Fig. 1.
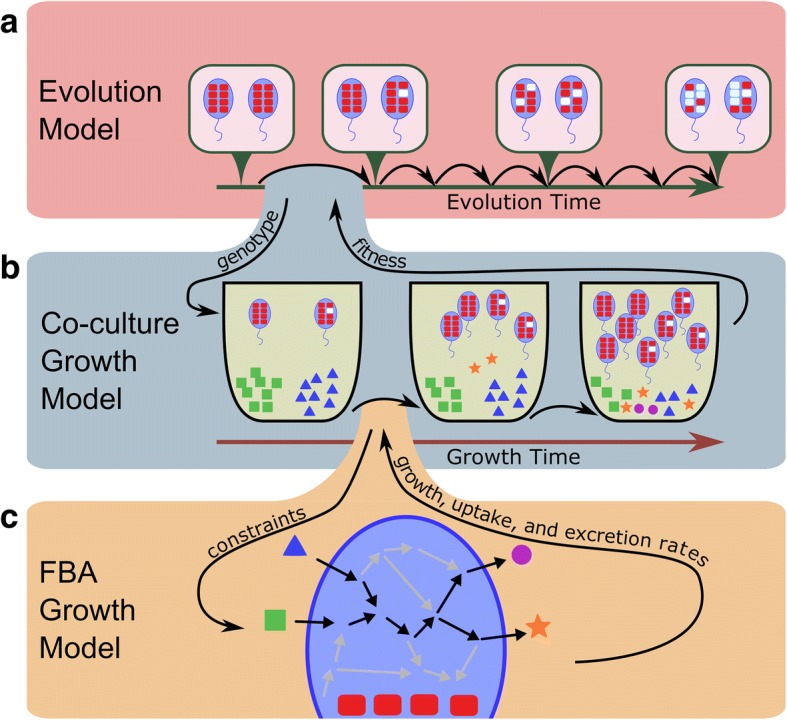
A framework for modeling the evolution of species interaction. a To model reductive evolution, genes are iteratively chosen at random as candidates for deletion, the fitness effect of their deletion is evaluated (using a co-culture growth model; panel (b)), and if the fitness effect is relatively small, these genes are deleted. b The co-culture growth model simulates the growth of the two species in a shared environment, and is based on a previously introduced dynamic multi-species model [28]. This model iteratively infers the behavior of each species in the shared environment based on an FBA approach (panel (c)). The predicted growth of each species and the predicted rates at which it uptakes and excretes various metabolites are used to update the abundances of species in the co-culture and the concentration of metabolites in the shared environment over time. c An FBA model is used to predict the growth of each species in a given environment based on the set of metabolic reactions and constraints encoded by the species and the concentration of metabolites in its environment
