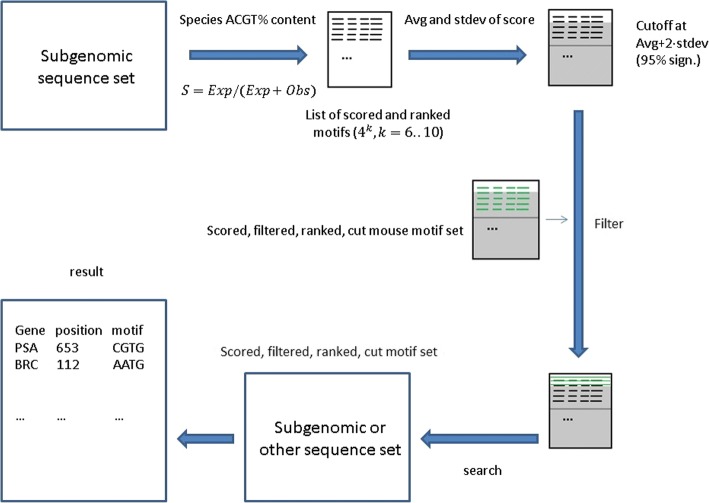
Fig. 1.
Graphical overview of application of algorithm. First, the A, C, G, T% for a given subgenomic sequence set is determined. Next, the statistics for each motif (for a given motif length from 6 to 10) is generated. This involves the calculation of the actual and expected occurrence for each motif as well as the score for each motif. Next, the set of motifs are filtered whose score is at least two standard deviations above the average score value. Next, the motif set is filtered again to remove general mammalian motifs. For this, the corresponding mouse motif set was used. Lastly, this set of statistically significant, double-filtered motif set can be used to search a given set of subgenomic or other sequence set