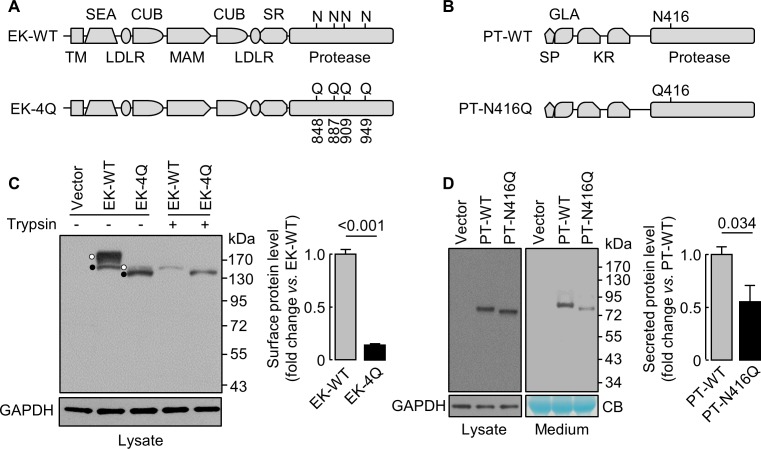
Figure 7. Analysis of N-glycosylation sites in the protease domain of EK and prothrombin.
(A) Illustration of EK WT and the mutant lacking the indicated N-glycosylation sites (EK-4Q). EK domains include transmembrane (TM), SEA, LDLR, CUB, MAM, scavenger receptor (SR) and protease domains. (B) Illustration of prothrombin WT and the PT-N416Q mutant lacking the N-glycosylation site in the protease domain. Prothrombin domains include signal peptide (SP), Gla (GLA), kringle (KR) and protease domains. (C) Western blotting of EK-WT and EK-4Q in HEK293 cells without (-) or with (+) trypsin treatment before the cells were lysed. The cell surface (trypsin-sensitive; white dots) and intracellular (trypsin-resistant; black dots) bands are indicated. Relative levels of surface EK bands in EK-WT and EK-4Q were estimated by densitometric analysis of western blots. Data are means ± S.E. from four independent experiments. p-Value is shown. (D) Western blotting of PT-WT and PT-N416Q in cell lysates (left) and the medium (right) from HEK293 cells. Relative levels of PT-WT and PT-N416Q in the medium were estimated by densitometric analysis of western blots. Levels of GAPDH in cell lysates and a Coomassie Blue (CB)-stained non-specific protein in the conditioned medium were used to assess protein amounts in each sample. Data are means ± S.E. from four independent experiments. p-Value is shown.