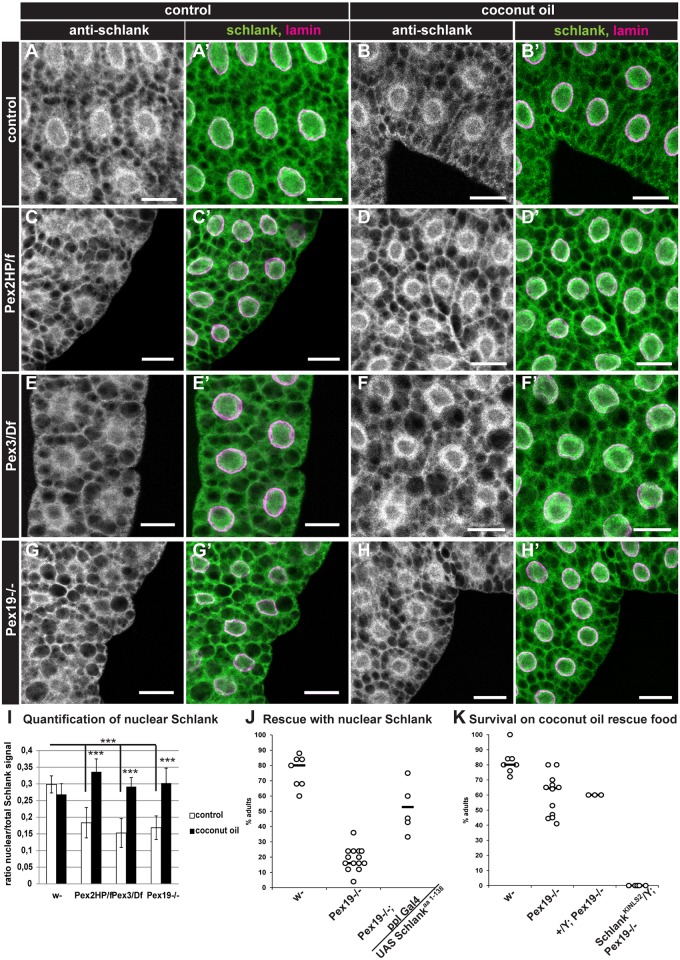
Fig 5. The dietary rescue functions by restoring Schlank nuclear localization.
(A-H) Staining of larval fat body tissue with α-Schlank and α-Lamin. Scale bars represent 20 μm. (I) Quantification of the Schlank nuclear fluorescent signal. Bars represent the ratio between CTCF from nuclear region (within Lamin staining) and whole cell. (J) Constitutively nuclear Schlankaa1–138 rescues Pex19 mutants. Genotypes are w-, Pex19ΔF7/Pex19ΔF7, Pex19ΔF7UAS Schlankaa1–138/Pex19ΔF7; ppl-Gal4/+. (K) Pex19 Schlank KINLS2 double mutants are not rescued by the M/LCFA-enriched diet. Genotypes are w-, Pex19ΔF7/Pex19ΔF7, +/Y; Pex19ΔF7/Pex19ΔF7, SchlankKINLS2/Y; Pex19ΔF7/Pex19ΔF7. n ≥ 3 in groups of 25 individuals. Error bars represent SD. Dots represent single experiments. Black bars represent median. *p > 0.05, ***p < 0.001. Significance tested using ANOVA with Tukey posttest. Corresponding raw data can be found in supplemental file S1 Data. aa, amino acid; CTCF, corrected total cell fluorescence; KINLS2, knock-in nuclear localization sequence 2; M/LCFA, medium- and long-chain fatty acid; UAS, upstream activating sequence.