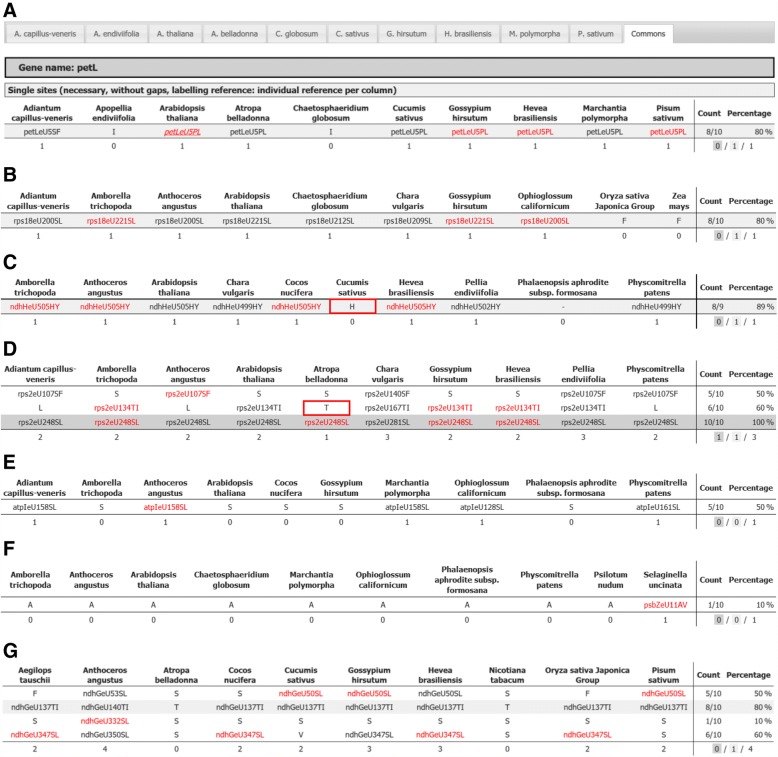
Fig. 2.
a-g Examples of the PREPACT 3.0 “commons” tab output for selected chloroplast queries as discussed in the text. For clarity of display, 10 of the now available 27 chloroplast references have been selected arbitrarily in each case. RNA editing prognoses are given in black when based on a “pre-edited” codon already present, but in red when based on a known RNA editing event in the respective organelle genome reference. The enhanced commons output now also displays amino acid identities for those references, which do not contribute to predict RNA editing events either because the unedited state is retained or because an inconvertible codon identity is present. The case of petL (a) and rps18 (b) are given as examples discussed in the text. The use of hyphens is now restricted to cases of lacking homology, such as the case of the ndh genes in Phalaenopsis (c). Documentation of RNA editing event ndhHeU505HY in Anthoceros and Hevea (c) supported that it was previously overlooked in Cucumis. Like the case of rps2eU134TI in Atropa (d), these candidate editing sites (red boxes) are now confirmed as previously overlooked RNA editing events (Additional file 2). The cases of evolutionary ancestral editing events rps2eU107SF (d) and atpIeU158SL (e) in the hornwort Anthoceros lacking in angiosperms suggest a shift of amino acid conservation making RNA editing obsolete. Rarely, yet other cases may reflect isolated “orphan” editing such as in Selaginella psbZ (f) or RNA editing that merely serves to alter overall hydrophobicity than affecting relevant individual codons like in ndhG (g)