Fig. 2.
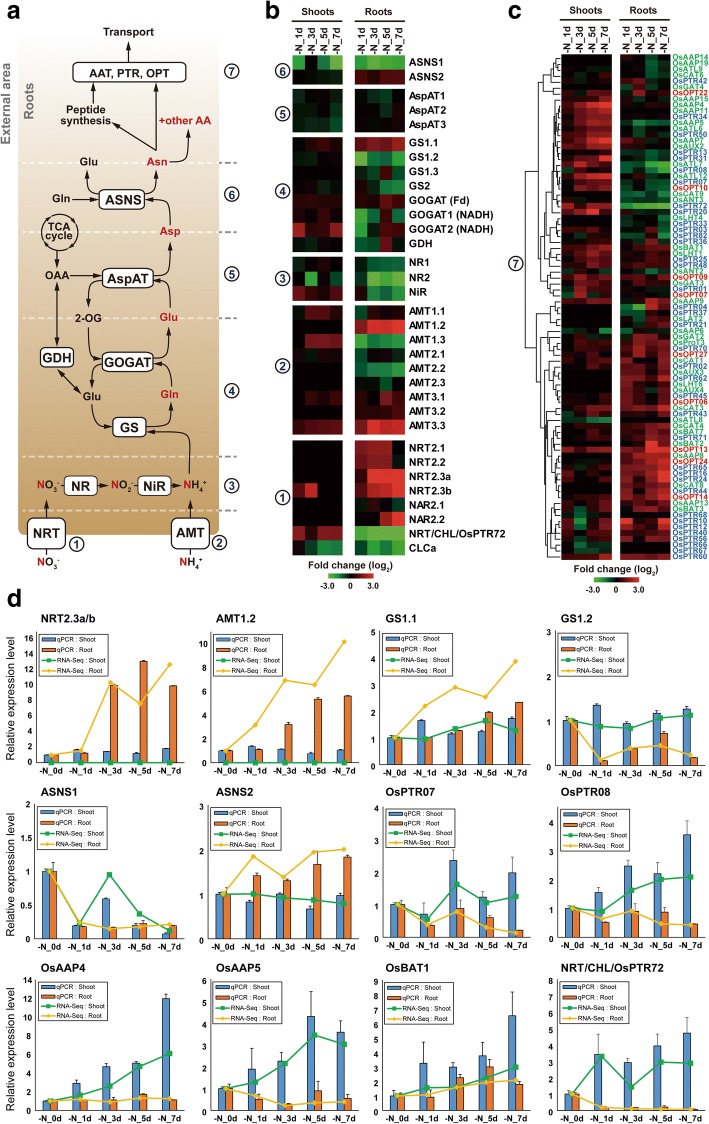
Expression profiles of genes involved in nitrogen (N) source uptake, assimilation, and transport in shoots and roots of N-starved rice. a Outline of N uptake, assimilation, and amino acid transport. (1) NRTs (nitrate transporters); (2) AMTs (ammonium transporters); (3) NRs (nitrate reductases) and NiR (nitrite reductase); (4) GSs (glutamine synthetases), GOGATs (glutamine-oxoglutarate aminotransferases), and GDH (glutamate dehydrogenase); (5) AspATs (aspartate aminotransferases); (6) ASNSs (asparagine synthetases); (7) AATs (amino acid transporters), PTRs (peptide transporters), and OPTs (oligopeptide transporters). Glu, glutamate; Gln, glutamine; OAA, oxaloacetate; 2-OG, 2-oxoglutarate; Asp, aspartate; Asn, asparagine. b Heatmap visualization of expression profiles of inorganic N source transporters (1–2) and assimilation-involved enzymes (3–6) in N-starved (-N_1d/3d/5d/7d) rice shoots and roots. c Expression profiles of 39 amino acid transporters (AATs), 37 peptide transporters (PTRs), and nine oligopeptide transporters (OPTs) in response to N starvation. Genes > 2-fold up- or down-regulated in at least one of four N-starved conditions (-N_1d/3d/5d/7d) in roots and/or shoots are shown. d Quantitative RT-PCR analysis of nitrate transporters (NRTs), ammonium transporters (AMTs), glutamine synthetases (GSs), asparagine synthetases (ASNSs), peptide transporters (PTRs), amino acid permeases (AAPs), and oligopeptide transporters (OPTs) in N-starved rice