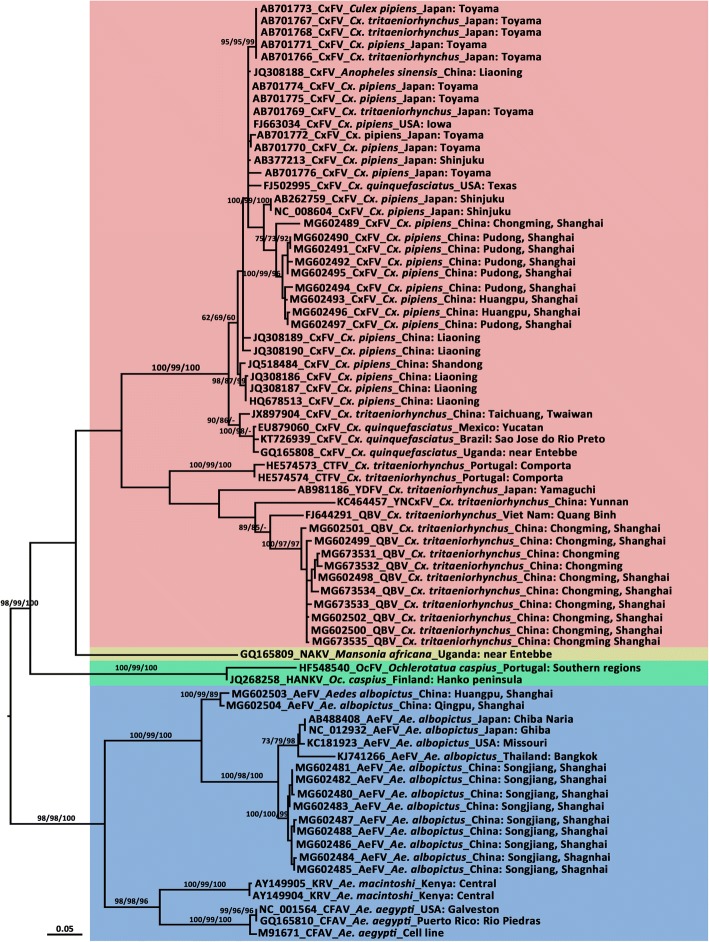
Fig. 1.
Molecular phylogeny of the 30 insect-specific flavivirus (ISFV) strains from Shanghai and other ISFVs based on partial NS5 gene (261 base pairs) sequences. The maximum likelihood tree was constructed by the GTR + I + R model. Sequences referable to the same host-genus are shown with the same colours in the tree. Bootstrap values (1000 replicates, not shown for less than 60%) of maximum likelihood, neighbour-joining, and Bayesian analyses are shown above the main lineages. The bar indicates 0.05 substitutions per site. AEFV, Aedes flavivirus; CFAV, cell fusing agent virus; CTFV, Culex theileri flavivirus; CxFV, Culex flavivirus; HANKV, Hanko virus; KRV, Kamiti River virus; NAKV, Nakiwogo virus; QBV, Quang Binh virus; OcFV, Ochlerotatua flavivirus; SCxFV, Spanish Culex flavivirus; YDFV, Yamadai flavivirus; YNCxFV, Yunnan Culex flavivirus. See Table 1 for information on the sequences of the Shanghai strains of ISFVs