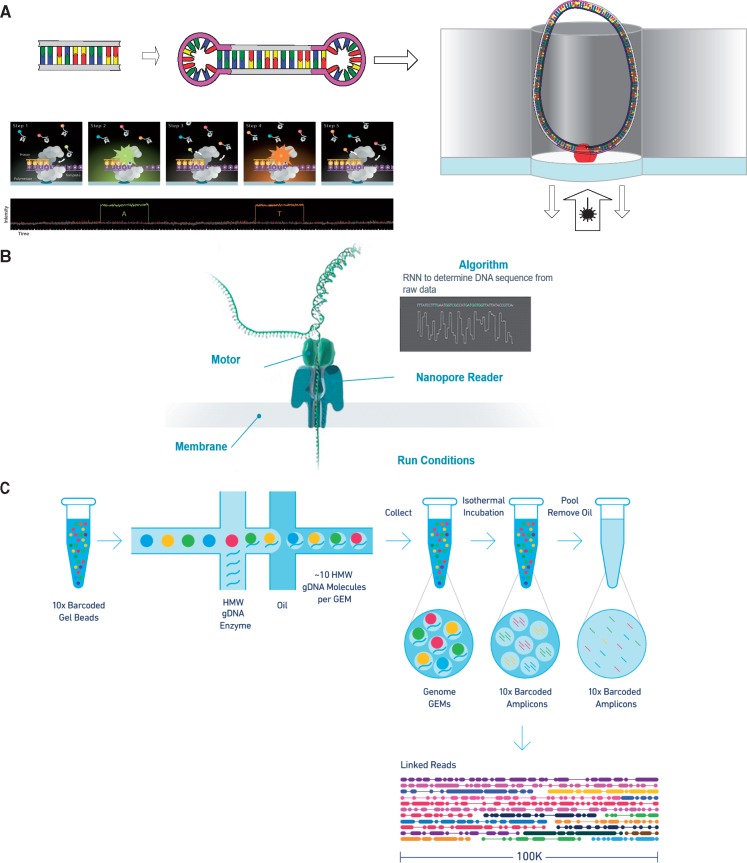
Figure 2.
Long-read sequencing technologies. (A) PacBio SMRT sequencing. Double stranded DNA is first sheared and size selected to the desired length and then sequencing adaptors are annealed. The adaptors are bound to a sequencing primer and strand displacing polymerase which adheres to the bottom of a well containing a zero mode wave guide. Following a pre-extension period where the polymerase reaction is run in the dark, the fragment is illuminated with a laser and as each base in the sequencing solution is incorporated, the fluorophore is detected and the polymerase reaction displaces it, giving a time and intensity signal which is converted into a base call. (B) Oxford Nanopore Technology passes the DNA molecule through a nanopore attached the flow cell surface membrane. As each base of the DNA molecule passes through the pore changes to the current passing through the pore are detected and converted into a signal. The signal detected is passed to a recurrent neural network (RNN) which converts it into base calls. (C) 10X Genomics Chromium technology works by means of an emulsion droplet technology, where gel beads are mixed with high molecular weight genomic DNA and an enzyme. Within each gel bead DNA is sheared and barcoded, creating fragments which can then be sequenced with Illumina sequencing. The presence of the chromium barcode then provides a mapper or assembler with linked-reads, allowing the relative spatial position of the fragments to be estimated Components of figure reproduced with permission from Pacific Biosciences, Oxford Nanopore Technologies and 10X Genomics.