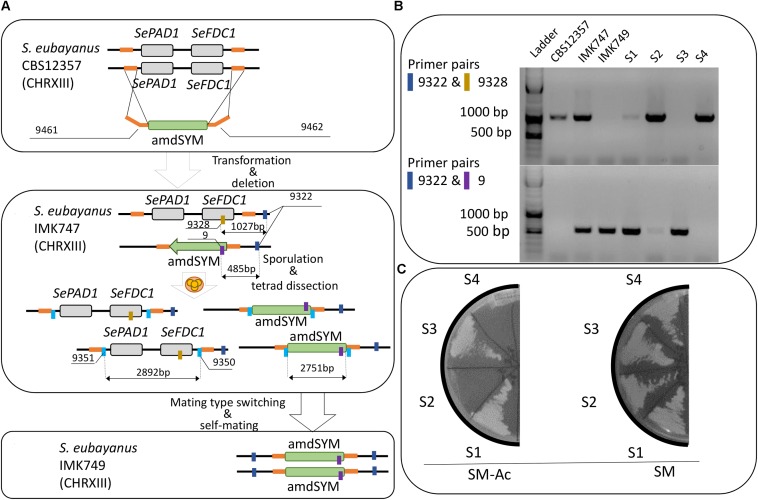
FIGURE 1.
Construction of targeted PAD1-FDC1 deletion in Saccharomyces eubayanus CBS 12357. (A-1) The deletion cassette was amplified using the primers pair 3144/3145 and the plasmid pUG-amdSYM (Solis-Escalante et al., 2013) as template. The fragment was then transformed in S. eubayanus CBS 12357. (2) Transformants were genotyped with primers pairs 9322/9328 and 9322/9 to discriminate between wild type (1027 bp) and deleted (485 bp) loci respectively. Heterozygote diploid IMK747 (SePAD1-SeFDC1/Sepad1-Sefdc1Δ::amdSYM) was sporulated. After ascus digestion the spores were dissected resulting in a two:two segregation of the PAD1-FDC1 locus. Spores were germinated. Single colony isolates of acetamide growing spores were selected and checked for ploidy. (3) A 2n isolated homozygote (Sepad1-Sefdc1Δ::amdSYM/Sepad1-Sefdc1Δ::amdSYM) was selected and renamed IMK749. (B) PCR analysis was performed to confirm correct integration of the gene disruption cassettes using amdSYM (Solis-Escalante et al., 2013) of the PAD1-FDC1 locus in S. eubayanus. PCR was carried out on reference strain CBS 12357, IMK747, IMX749 and four segregants (S1–S4) of a single tetrad of IMK747. PCRs were performed with the primers pairs 9322/9328 and 9322/9 for PAD1-FDC1 and pad1-fdc1Δ loci, respectively. The amplification of the wild-type PAD1-FDC1 and pad1-fdc1Δ parental strain, of the loci generated fragments of 1027 and 485 bp, respectively. (C) The four segregants (S1–S4) of a single tetrad of IMK747 were grown on SM-Ac and SM. The plates were incubated at 30°C and were read after 3 days.