Fig. 4.
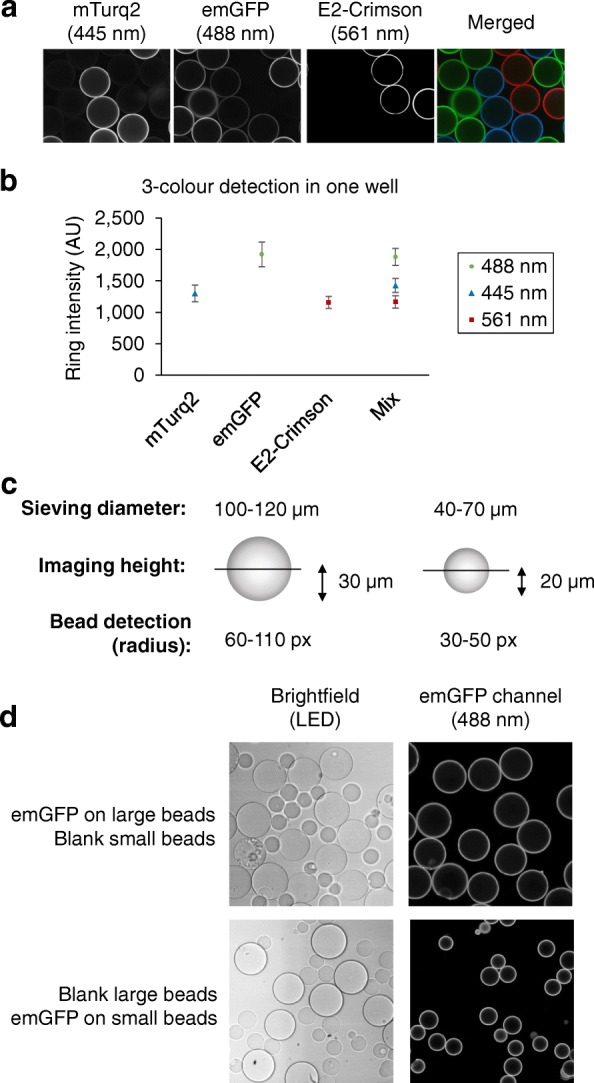
Detection of mixed bead populations by different color or by bead size. a, b Three bead-bound fluorescent proteins detected in a single well. a Three different His6-tagged fluorescent proteins (mTurquoise2 (mTurq2, blue), emGFP (green) and E2-Crimson (red)) were pre-immobilized on individual batches of Ni2+NTA agarose beads. The beads were then pooled together in one well of a 384-well plate. Images were acquired using the confocal scanning microscope Opera™ (Perkin Elmer) with 445, 488, and 561 nm excitation wavelengths, and detected in three emission channels (475/34, 520/35, 660/150, and 690/70 nm, detailed in Additional file 1: Figure S3). b Images of wells with single and mixed bead populations of mTurquoise2, emGFP, and E2-Crimson beads were analyzed and average ring intensities were calculated for each channel. c Differential detection of bead populations by size. Large beads were sieved through pore size mesh of 100 and 120 μm, imaged on the Opera™ (Perkin Elmer) at a confocal plane height of 30 μm, and identified in the 60–110 pixel radius range. Small beads were sieved through pore size filters of 40 and 70 μm, imaged on the Opera at the confocal plane of 20 μm and detected in the 30–50 pixel radius range. d Each bead is a separate reaction unit. His6-tagged emGFP was immobilized on large or small beads and mixed in one well with blank small or blank large beads. Beads were detected as specified in (c) in brightfield and emGFP fluorescence emission channels. No emGFP was detected on the blank beads present in the same well, indicating the absence of protein transfer between the beads
