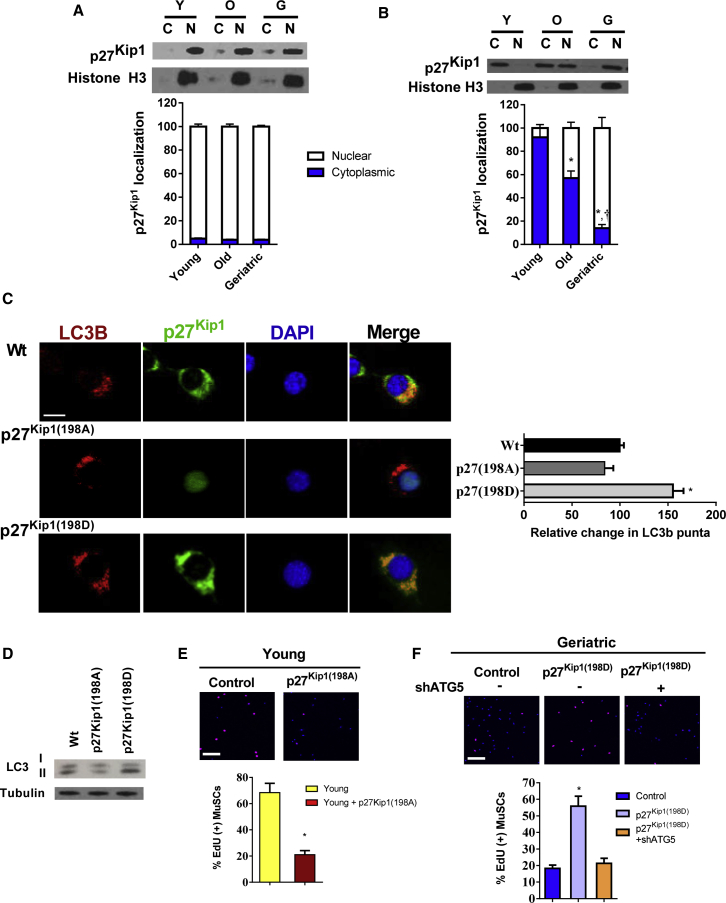
Figure 5.
Cellular Location of p27Kip1 Regulates MuSC Function in Part through the Induction of Autophagy
(A) Upper: p27Kip1 protein expression in both cytoplasmic and nuclear compartments of freshly sorted MuSCs across age groups. Lower: quantification of percentage of cytoplasmic and nuclear expression of p27Kip1 (N = 3 independent experiments).
(B) Upper: p27Kip1 protein expression in both cytoplasmic and nuclear compartments of activated MuSCs (48 hr in culture) across age groups. Lower: quantification of percentage of cytoplasmic and nuclear expression of p27Kip1 (N = 3 independent experiments).
(C) p27Kip1 phosphorylation status-mediated cellular location and autophagy. Left: immunofluorescence staining of LC3B and p27Kip1 expression of young MuSCs overexpressing GPF-labeled WT p27Kip1, p27Kip1(198A), or p27Kip1(198D). All cells were treated with chloroquine for the final 12 hr of culture. LC3B, red; p27Kip1, green; DAPI, blue. Scale bar, 5 μm in (C). Right: quantification of relative change in LC3B puncta across treatment groups (N = 3 independent experiments).
(D) LC3B isoform protein expression in young MuSCs overexpressing WT p27Kip1, p27Kip1(198A), or p27Kip1(198D).
(E) Upper: EdU incorporation in control or p27Kip1(198A) overexpressing young MuSCs. Lower: quantification of percentage of EdU(+) MuSCs in each group (N = 3 independent experiments).
(F) Upper: EdU incorporation in control, p27Kip1(198A) overexpressing or p27Kip1(198D) overexpressing geriatric MuSCs with or without Atg5 knockdown. Lower: quantification of percentage of EdU(+) MuSCs in each group (N = 3 independent experiments).
∗ Signifies difference from young or control cells; † signifies difference from old cells (p < 0.05). Scale bars, 100 μm in (E) and (F). Data are presented as means ± SE.