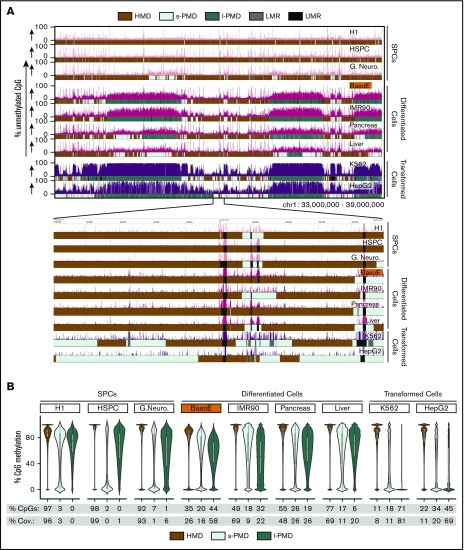
Figure 1.
Diversity of global methylation patterns in HSPC, differentiated, and transformed cells. (A) The percentage of unmethylated CpG dinucleotides is plotted for a typical region on chr1. Genplay screenshot of a 6-megabase region on chromosome 1 that illustrates the Methyl-Seq data and the segmentation of the methylome. The inset below the main panel represents the indicated 200-kb region at the center of the 6-megabase region to illustrate the fine structure of the UMRs and LMRs. The colored boxes below each track illustrate the cell-specific segmentation of the genome into HMDs, s-PMDs, l-PMDs, LMRs, and UMRs. The methylome of SPCs is composed mostly of HMDs and of small unmethylated regions (UMRs and LMRs) that correspond to regulatory elements. Methylomes of differentiated and transformed cells also contain HMDs, UMRs, and LMRs, but PMDs represent a large fraction of the genome of these cells. PMDs are ∼30% to 70% methylated in differentiated cells but are almost completely unmethylated in transformed cells. (B) Violin plots illustrating DNA methylation density in HMDs and PMDs. The percentages of CpGs and coverage (% Cov.) of the genome that are in HMDs, l-PMDs, and s-PMDs are indicated below the plots. SPCs are composed mostly of HMDs; differentiated cells contain variable amounts of HMDs and PMDs while transformed cells contain small amounts of HMDs.