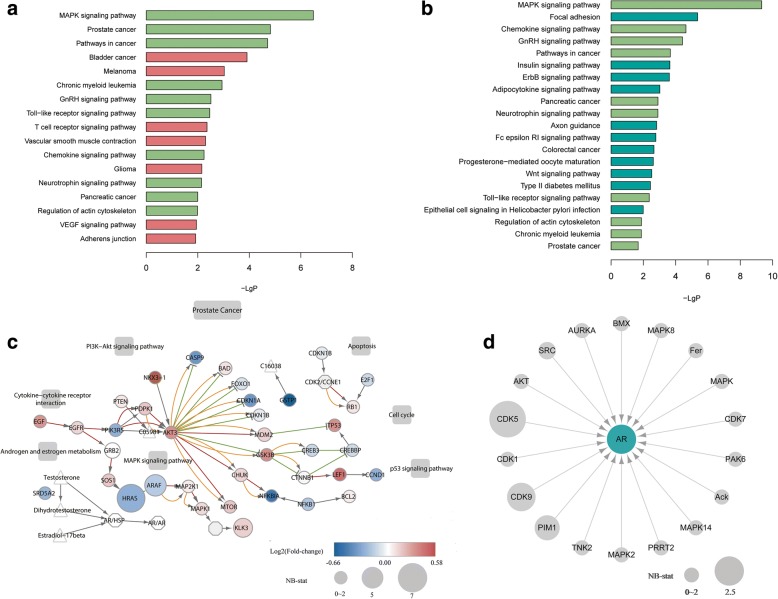
Fig. 3.
Functional enrichment analysis of DE and DS kinase genes. a Enriched KEGG pathways of DE kinase genes. b Enriched KEGG pathways of DS kinase genes. Pathways enriched in both DE and DS kinase genes are shown in light green in both plots. Those only enriched in DE kinase genes are shown in purple in (a), and those only enriched in DS kinase genes are shown in dark green in (b). c The “Prostate Cancer” pathway in KEGG, with detected DE and DS genes highlighted in colors. Circles represent genes, with their colors showing their fold-change between the cancer and normal groups. Circle size indicates the NB-stat calculated by DSGseq, which represents the degree of differential splicing of the gene between the two groups. Small molecules (triangles), cellular process (rectangles) and gene complexes (hexagon) are also shown. Colored edges indicate activation (red), inhibition (green), phosphorylation (orange). d Expression and splicing changes of kinases that phosphorylate AR. None of the kinase genes is differentially expressed, but CDK5, CDK9, PIM1 and SRC are differentially spliced, with node sizes representing the NB-stat in DSGseq