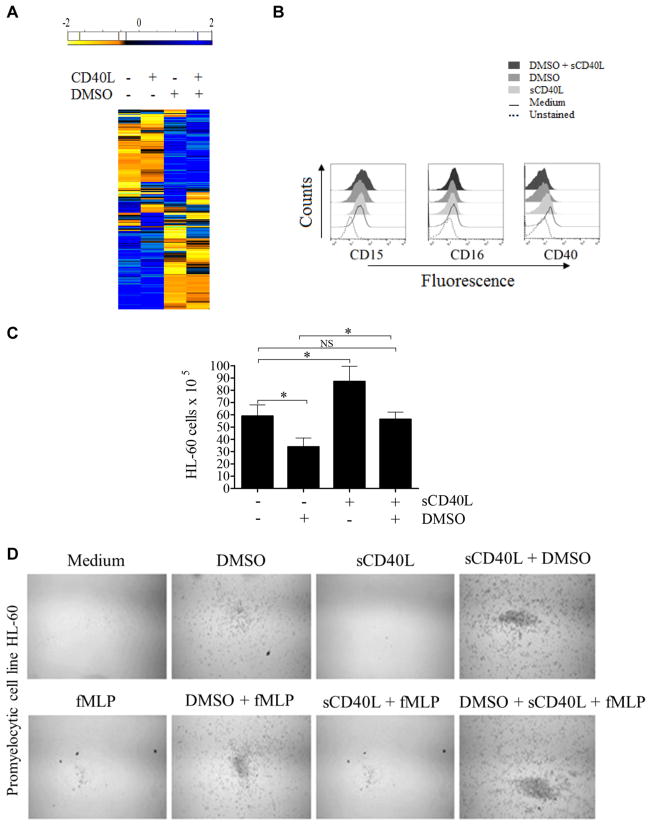
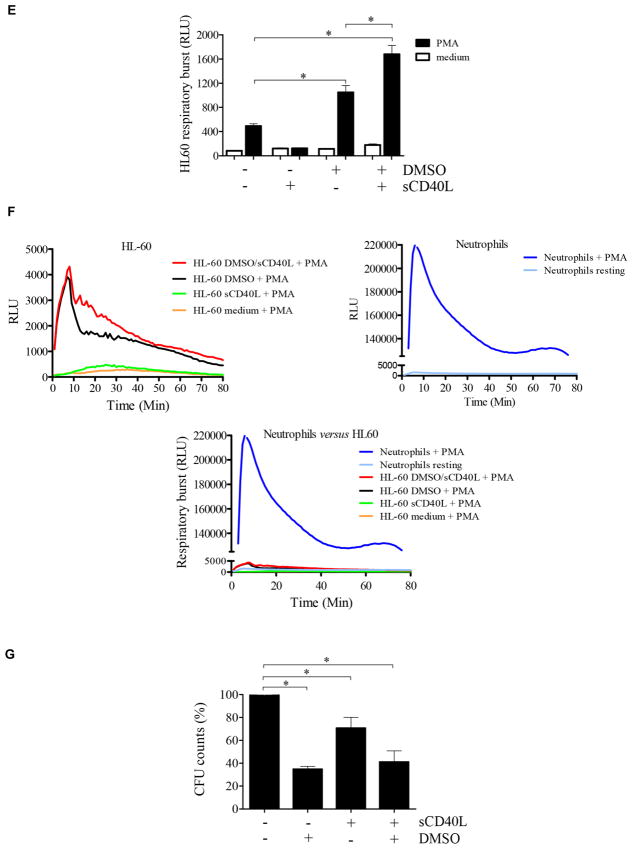
FIG 5.
sCD40L regulates promyelocytic cell development. A, Heat map showing that sCD40L induces transcriptome alterations in the promyelocytic HL-60 cells. HL-60 cells were incubated for 48 hours in the absence and presence of 500 ng/mL sCD40L, 1% DMSO, or both before RNAseq analysis. In the heat map the results obtained in transcripts per million (TPM) are represented in a Log2 scale, where yellow shows low expression and blue shows high expression. Gene Ontology analysis of genes upregulated or downregulated by sCD40L compared with untreated cells or DMSO-treated versus DMSO plus sCD40L–treated cell are listed in Table I. A pool of 3 independent cultures of HL-60 cells in which every experimental condition (resting, DMSO, sCD40L, and DMSO plus sCD40L) has been performed in triplicates. B, Histograms obtained by using flow cytometric analysis showing expression of CD15, CD16, and CD40 on HL-60 cells. C, Equal numbers of promyelocytic HL-60 cells were distributed in 6-well plates and cultured in the presence or absence of 1% DMSO, 500 ng/mL sCD40L, or both. After 6 days, cells were harvested and counted, and cell viability was assessed (see Fig E4). D, HL-60 migration was assessed by using a transwell migration assay. Microscopic images display cells that migrated. Data shown are representative of 3 independent experiments performed in triplicates. E, Respiratory burst of HL-60 cells was induced by PMA (90 mmol/L) and analyzed with a luminol-enhanced chemiluminescence assay. Values are expressed as relative light units (RLU). F, Comparative graphics showing the kinetics of respiratory burst of HL-60 cells (upper left panel), neutrophils (upper right panel), and both overlapping (lower panel). G, HL-60 cells were challenged with P brasiliensis (ratio of 2 neutrophils/1 fungus), and their microbicidal activity was assessed by counting CFU values from recovering internalized fungi. CFU values (in percentages) were determined in relation to CFU numbers of untreated HL-60 cells. *P ≤ .05. NS, Not significant.