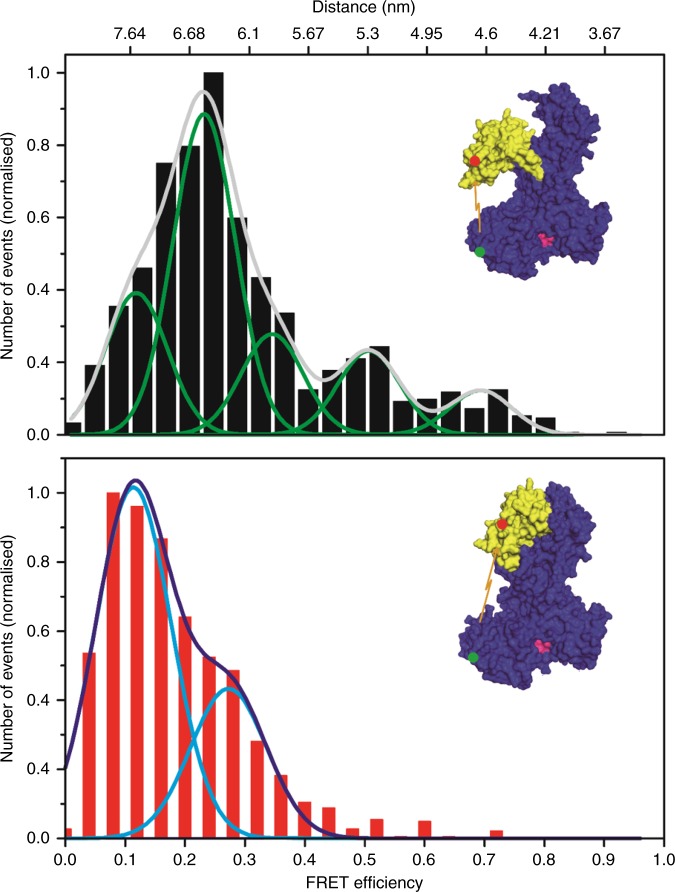
Fig. 3.
Upon adenosine triphosphate (ATP) binding, the pre-protein binding domain (PBD) moves away from the IRA2 domain. Histogram of the single-molecule Förster resonance energy transfer (smFRET) data obtained with SecA 329C–472C, in the absence (top) and the presence of ATP (bottom). While the data in the presence of ATP can be fitted with a bimodal Gaussian distribution, the data in the absence of nucleotide is best fitted with a multimodal function. The distance between the two fluorophores in the PBD and the IRA2 domain increases when ATP is present from 6.5 nm to 7.5 nm, indicating that the PDB moves towards the helical wing domain (HWD). Insets show the position of the fluorescent labels in SecA. To illustrate the movement of the PBD, the atomic SecA structures 1TF2 (top) and 1M74 (bottom) are shown. The distance between the PBD at residue 329 (red dot) and the IRA2 domain at 472 (green dot) is highlighted with arrows and is 5.1 nm in the B. subtilis open structure (1TF2) and 6.1 nm in the wide open structure (1M74) between the corresponding residues (D309 and K452)