Fig. 1.
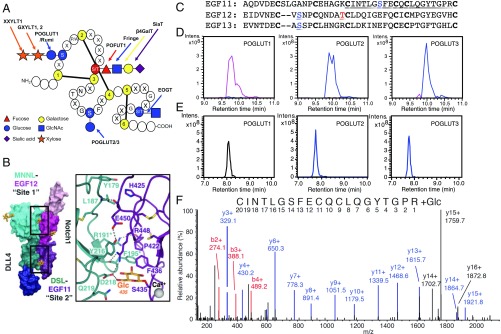
KDELC1 and KDELC2 are protein O-glucosyltransferases (POGLUTs). (A) Novel hexose exists at a distinct site from previously described O-Glc, O-Fuc, or O-GlcNAc modifications. Adapted from ref. 39. Copyright (2011), with permission from Elsevier. (B) Recent cocrystal structure of NOTCH1-DLL4 reveals a hexose modification on S435 of NOTCH1 EGF11 in contact with DLL4. From ref. 22. Reprinted with permission from AAAS. (C) Amino acid sequence of human NOTCH1 EGF11–13 with each EGF aligned based on conserved cysteines (boldface type). Known O-Glc and O-Fuc sites in EGF12 and -13 are indicated in blue and red, respectively. The O-Glc site in EGF11 (S435) is in blue, and the tryptic peptide containing this site described in E and F is underlined. (D) EICs of the reaction products of POGLUT1, POGLUT2, or POGLUT3 after overnight incubation with bacterially expressed human NOTCH1 EGF11–13 and UDP-Glc. Black line, unmodified EGF11–13 (+12, 1,236.5 m/z); blue line, EGF11–13 + 1 hexose (+12, 1,250.0 m/z); pink line, EGF11–13 + 2 hexoses (+12, 1,263.5 m/z). (E) The products from D were reduced/alkylated, digested with trypsin, and analyzed by LC-MS/MS. EICs were generated for different glycoforms of the triply charged peptide from EGF11 underlined in C. Black line, unmodified peptide (788.2 m/z). Blue line, peptide + 1 hexose (842.2 m/z). (F) HCD-MS/MS spectra of the O-glucosylated peptide from EGF11 confirms that S435 (equivalent to y-14 ion) is modified with O-Glc. Numerous fragment b-ions (red) and y-ions (blue) were detected. Note that y-ions 14 and 15 correspond to the peptide retaining a hexose (blue) although y-ions 14, 15, and 16 corresponding to the peptide ions that lost the hexose during fragmentation were also detected (black).