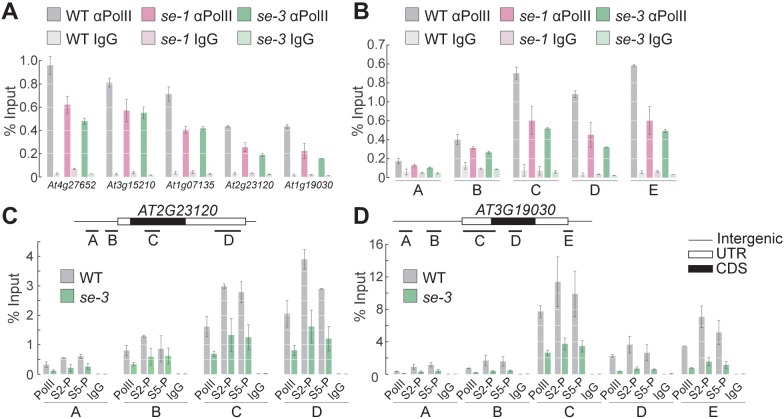
Figure 3. SE ensures efficient pol II association to intronless genes.
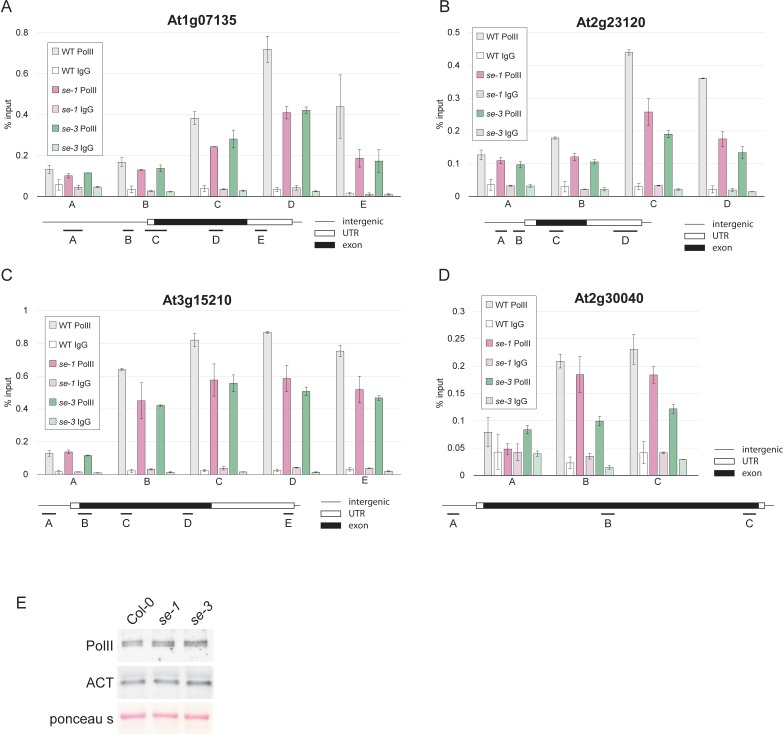
(A,B) Analysis of pol II occupancy at SE target loci by pol II-ChIP qPCR in WT, se-1 and se-3 in gene bodies of five different genes (A) or at various genomic regions at At3g19030 (B). Regions tested are depicted in panel D. A general pol II CTD antibody and mouse IgGs (as negative control) were used for immunoprecipitation. Additional gene loci were tested and are shown in Figure 3—figure supplement 1. Error bars indicate the range of two independent biological experiments. (C,D) Analysis of Ser5P and Ser2P pol II levels at SE target loci by ChIP qPCR in WT and se-3. General pol II CTD, pol II CTD Ser2P and pol II CTD Ser5P specific antibodies and mouse IgGs (as negative control) were used for immunoprecipitation. Additional gene loci were tested and are shown in Figure 3—figure supplement 3. Error bars indicate the range of two independent biological experiments.