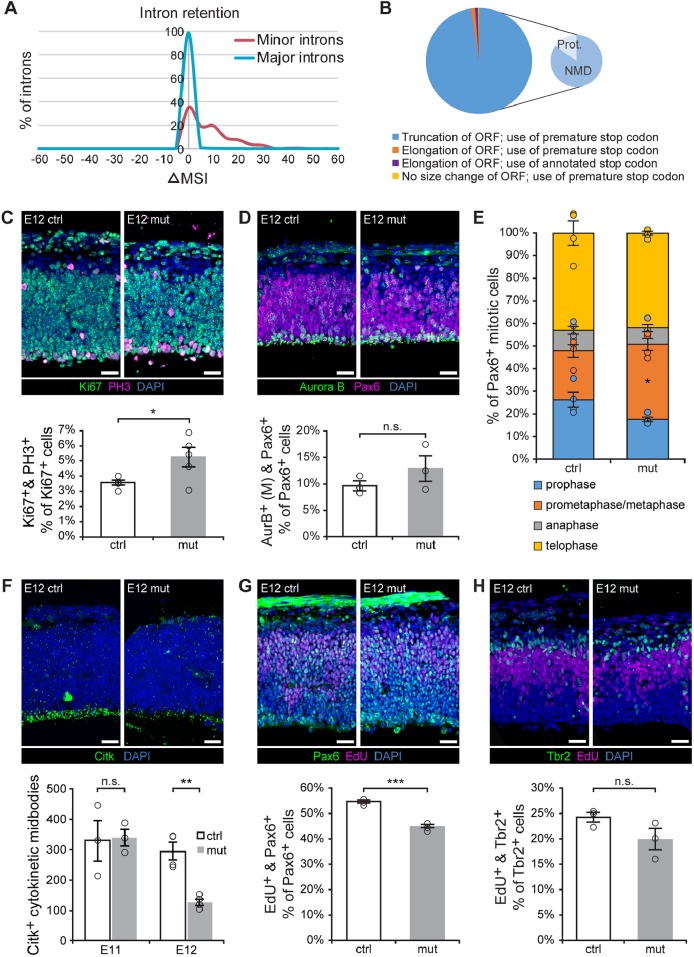
Fig. 5.
U11-null RGCs exhibit S-phase, metaphase and cytokinesis defects. (A) Frequency plot of the change in mis-splicing index (ΔMSI, x-axis) of major (blue) and minor (red) introns. (B) Pie chart showing the effect of minor intron retention on the open reading frame (ORF), for the 186 minor introns with significantly elevated retention, across all annotated MIG transcripts requiring minor intron splicing. The truncation category is further broken down into the percentage of transcripts predicted to be degraded by nonsense-mediated decay (NMD) or to produce protein (prot.). (C) IF for Ki67 (green) and PH3 (magenta) in sagittal sections of the E12 control (ctrl) and mutant (mut) pallium, with quantification. (D) IF for Aurora B (green, AurB) and Pax6 (magenta) in sagittal sections from E12 ctrl and mut pallium, with quantification of mitosis-specific Aurora B staining (M). (E) Quantification of the percentage of Aurora B+ cells in different mitotic phases for E12 ctrl and mut pallium. (F) IF for Citk (green) in sagittal sections of ctrl and mut E12 pallium, quantification of Citk+ cytokinetic midbodies along the ventricular edge in the E11 and E12 control and mutant pallium. (G) IF for Pax6 (green) and EdU (magenta) on E12 ctrl and mut sagittal pallial sections, with quantification. (H) IF for Tbr2 (green) and EdU (magenta) on sagittal sections of E12 ctrl and mut pallium, with quantification. Scale bars: 30 µm. Data are presented as mean±s.e.m. For details of statistical methods, see Table S8. n.s., not significant, *P<0.05, **P<0.01, ***P<0.001. See also Figs S3 and S4, and Tables S3 and S4.