Figure 2.
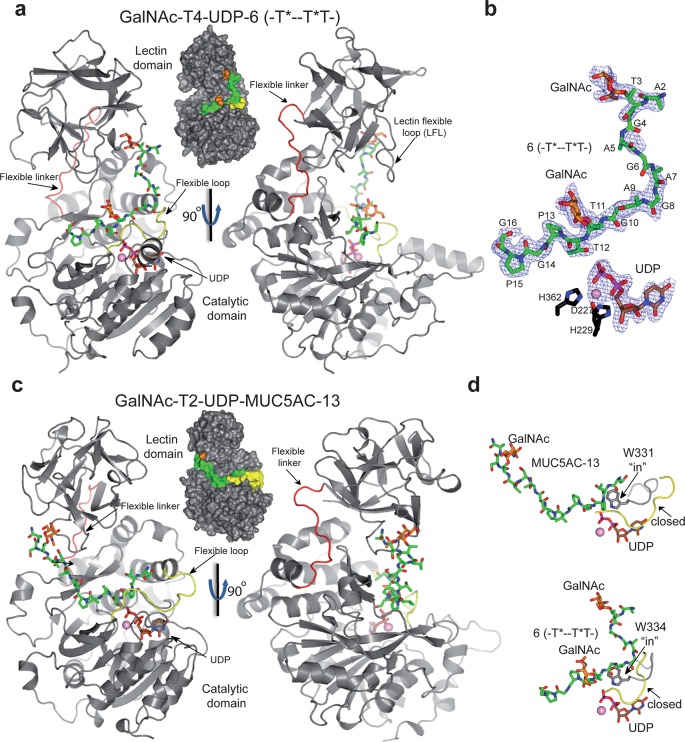
Crystal structure of GalNAc-T4 in complex with UDP-Mn+2 and glycopeptide 6. (a) Two different views of GalNAc-T4 in complex with 6. The catalytic and lectin domains are colored in gray, and the flexible linker and catalytic domain active site loop are depicted in red and yellow, respectively. The lectin domain flexible loop (LFL) is indicated by a black arrow. The GalNAc moiety of the Thr3-GalNAc and Thr11-GalNAc is shown as orange carbon atoms while the rest of the peptide is shown as green carbon atoms. The nucleotide is depicted as brown carbon atoms whereas the manganese atom is shown as a pink sphere. Inserted between the structures is a surface representation of GalNAc-T4 with the same orientation as the cartoon representation of the leftmost structure. (b) Electron density maps are FO–FC (blue) contoured at 2.0 σ for glycopeptide 6, UDP, and manganese ion. (c) Two different views of GalNAc-T2 in complex with MUC5AC-13 (PDB entry 5AJP(14)) again with an inserted surface representation between structures. Atom colors are the same as in part a above. (d) Close-up views of the GalNAc-T2-UDP-MUC5AC-13 and the GalNAc-T4-UDP-glycopeptide 6 complexes showing the bound glycopeptide and the catalytic domain active site flexible loop (in black). Note that flexible loop residues Trp331 in GalNAc-T2 and Trp334 in GalNAc-T4 adopt an “in” loop conformation.