Fig. 1: Overview and examples of gene entries in the Eurexpress, Gene Expression Atlas, and OMIM databases.
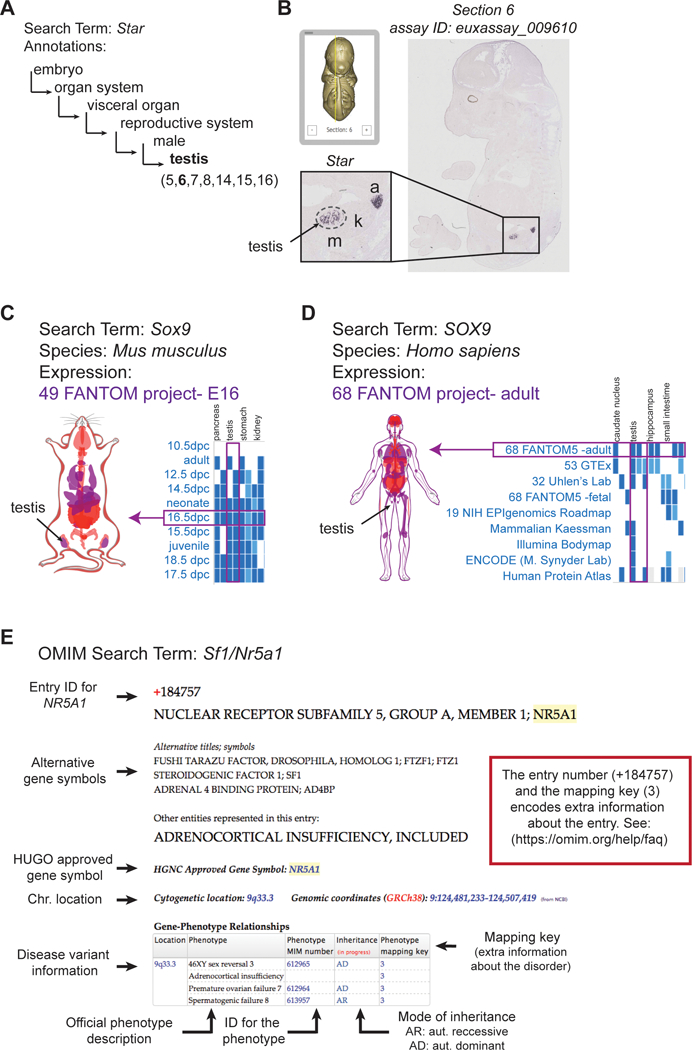
(A) In situ hybridization images of 14.5 dpc embryos are compiled in the Eurexpress Transcriptome Atlas Database for Mouse Embryo (http://www.eurexpress.org). Expression observed in an image is annotated and encoded as text that can be queried. The more general anatomical terms branch into more specific terms. For example, the nested terms that describe the testis are displayed. (B) The Eurexpress viewer scrolls through sections in a “virtual embryo environment” and zoom into regions as similar to a digital microscope. For the gene Star, the testis can be visualized in the 6th section. The testis region is demarcated by a dashed line; a, adrenal; k, kidney; m, mesonephros. (C, D) The Gene Expression Atlas houses easy to access transcriptomic data for a wide variety of species from all major consortia and sequencing efforts. For the gene Sox9, detailed information are available on expression in the (C) mouse and (D) human. Data is presented in a tabular format that can be sorted using a number of parameters (graphics shown here are part of the full table). Selecting data from a project in the table, such as 16.5 dpc 49 FANTOM (C, in mouse) or 68 FANTOM (D, in human), highlights the organs (in our case, the testis) that are included in that dataset in purple on the interactive body-map. (E) The OMIM database (Online Mendelian Inheritance in Man) catalogues human disease variants. Entries can be either by phenotype or gene. In the example of a search for the gene NR5A1 (also known as SF1), there are many alternative gene symbols that have been used historically; however, the disease information is catalogued under the HUGO approved symbol (NR5A1). Information on the location of the gene, all phenotypes caused by mutations in NR5A1 are listed in a table with ID numbers for each phenotype and additional information on inheritance. The entry number and mapping key encode additional information about the disorder.
