Abstract
Background
In spite of extensive study, the mechanisms for salt sensitivity of BP in humans and rodent models remain poorly understood. Several microRNAs (miRNAs) have been associated with hypertension, but few have been shown to contribute to its development.
Methods
We examined miRNA expression profiles in human kidney biopsy samples and rat models using small RNA deep sequencing. To inhibit an miRNA specifically in the kidney in conscious, freely moving rats, we placed indwelling catheters to allow both renal interstitial administration of a specific anti-miR and measurement of BP. A rat with heterozygous disruption of the gene encoding endothelial nitric oxide synthase (eNOS) was developed. We used bioinformatic analysis to evaluate the relationship between 283 BP-associated human single-nucleotide polymorphisms (SNPs) and 1870 human miRNA precursors, as well as other molecular and cellular methods.
Results
Compared with salt-insensitive SS.13BN26 rats, Dahl salt-sensitive (SS) rats showed an upregulation of miR-214-3p, encoded by a gene in the SS.13BN26 congenic region. Kidney-specific inhibition of miR-214-3p significantly attenuated salt-induced hypertension and albuminuria in SS rats. miR-214-3p directly targeted eNOS. The effect of miR-214-3p inhibition on hypertension and albuminuria was abrogated in SS rats with heterozygous loss of eNOS. Human kidney biopsy specimens from patients with hypertension or hypertensive nephrosclerosis showed upregulation of miR-214-3p; the gene encoding miR-214-3p was one of several differentially expressed miRNA genes located in proximity to human BP-associated SNPs.
Conclusions
Renal miR-214-3p plays a functional and potentially genetic role in the development of hypertension, which might be mediated in part by targeting eNOS.
Keywords: hypertension, kidney, microRNA, nitric oxide, genetics
Visual Abstract
Hypertension is the leading risk factor for death worldwide.1 Genome-wide association studies (GWASs) have identified several hundred single-nucleotide polymorphisms (SNPs) associated with BP.2–6 Exacerbated BP salt sensitivity is present in the majority of patients with hypertension.7 The kidney plays an essential physiologic role in BP regulation and the development of hypertension including salt-sensitive hypertension.8,9 However, the molecular and mechanistic basis for hypertension and BP salt sensitivity remains poorly understood.
microRNAs (miRNAs) participate in the regulation of a wide range of cellular functions primarily by binding to the 3′ untranslated region (3′UTR) of target mRNAs and decreasing the abundance of target mRNAs or proteins. Several miRNAs are differentially expressed in patients with hypertension or animal models of hypertension.10,11 Dozens of miRNAs regulate protein-coding genes relevant to BP regulation.12 However, very few miRNAs have been shown to be genetically or functionally involved in the development of hypertension.13
The Dahl salt-sensitive (SS) rat is the most widely used polygenic model of human salt-sensitive hypertension.7 Several physiologic pathways, including pathways in the kidney, may contribute to the development of hypertension in SS rats.9,14 The genetic cause of the exacerbated salt sensitivity in the SS rat remains unclear.15 Replacement of a 13-million-base-pair region of chromosome 13 in the SS rat genome with the corresponding chromosomal segment from the salt-resistant Brown Norway (BN) rat, which results in a congenic rat strain named SS.13BN26, leads to significant attenuation of salt-sensitive hypertension and renal injury.16,17 It indicates that the SS.13BN26 congenic region harbors genes that influence BP salt sensitivity.
The goal of this study was to examine whether any specific miRNAs in the kidney contribute to the development of hypertension and identify the underlying mechanism. A secondary goal of the study was to examine whether any miRNA was involved in the genetic regulation of BP or hypertension. We used a wide range of approaches including bioinformatic analysis of human GWAS data, small RNA deep sequencing analysis of scarcely available human kidney biopsy specimens, kidney-specific inhibition of an miRNA, and a newly generated rat strain with a targeted gene disruption. The results indicate that miR-214-3p in the kidney contributes to salt-induced hypertension functionally and potentially genetically. The study also identified miR-214-3p as a direct regulator of endothelial nitric oxide synthase (eNOS), which mediates, at least in part, the effect of miR-214-3p on hypertension.
Methods
Detailed methods are available in the Supplemental Material.
Animals
Dahl salt-sensitive SS/JrHsdMcwi (SS) rats and SS.BN-(D13Rat151-D13Rat197)/Mcwi (SS.13BN26) rats were developed or maintained as described previoulsy.16–18 All experimental procedures in this study involving animals were approved by the Institutional Animal Care and Use Committee of the Medical College of Wisconsin.
Small RNA Deep Sequencing and Data Analysis
RNA extraction and small RNA deep sequencing analysis were performed as described previously.12,19,20
Real-Time PCR
Measurement of miRNA abundance by real-time PCR was performed using Taqman assays (Applied Biosystems) as described previously.10,21
Renal Medullary Interstitial Infusion and Measurement of Arterial BP in Conscious, Freely Moving Rats
Rats were uninephrectomized to ensure that the remaining kidney receiving anti-miR injection was the sole determinant of kidney function. Catheterization to allow renal medullary interstitial delivery of anti-miR and measurement of arterial BP in conscious, freely moving rats was performed as described previously.22–25 Similar methods have been used by several laboratories for medulla-targeted, intrarenal delivery of compounds.26–30
Anti-miR or miRNA Mimic Oligonucleotides
Locked nucleic acid–modified anti-miR oligonucleotides (Exiqon) (2 mg/kg body wt) or miRNA mimics (67 nM) were used, similar to that described previously.10,31,32
3′UTR Reporter Assay
3′UTR reporter assay was performed in HeLa cells with pMIR-REPORT β-galactosidase as the normalizer as we described previously.10,32,33
Western Blot
Western blot was performed as described previously.21,34 Coomassie blue staining of the entire membrane was used for normalization.21,34
Generation of eNOS Gene Disruption on the Background of the SS Rat
eNOS gene (Nos3) disruption on the genetic background of the SS rat was produced using zinc-finger nucleases (ZFNs) targeting as described previously.17,35,36 Briefly, ZFNs targeting the Nos3 exon 3 sequence GACTTCATCAATCAGTACtataaCTCGATCAAAAGGTGGGT, where each ZFN binds to each underlined sequence on complementary strands, were obtained from Sigma-Aldrich. In vitro–transcribed mRNA encoding ZFNs was injected into SS/JrHsdMcwi (SS) strain embryos and a putative founder animal was identified harboring a 165-bp deletion mutation (Rn5, Chr13: 7350700–7350864). This founder was back-crossed to the parental SS strain and an SS-Nos3em13Mcwi (SS-Nos3+/−) breeding colony was established (Rat Genome Database ID: 6893431). For the described studies, SS-Nos3+/− animals were bred with parental wild-type SS animals and SS-Nos3+/− heterozygous and wild-type littermates were compared.
Bioinformatic Analysis of Human BP-Associated SNPs
Linux and R, including the R GenomicRanges package,37 were used to analyze the relationship between miRNAs and human BP-associated SNPs. The analysis was on the basis of hg38 genome assembly. SNPs associated with BP were retrieved from several recent GWASs.2–6 Redundant SNPs were removed, yielding a total of 283 SNPs. Chromosomal locations of the SNPs and associated genes were obtained from the NCBI DBSNP database. Chromosomal coordinates of linkage disequilibrium (LD) blocks for the SNPs were obtained using HaploReg v4.1 with default settings for LD blocks (r2=0.8) and matching ancestry.38 Information for human precursor microRNAs was obtained from miRBase.39 Genome coordinates (hg19) for 3′UTR regions of human coding genes were downloaded from TargetScan and converted to hg38 using the liftOver tool. The bed files of broadPeaks were downloaded from ENCODE. Data for enhancer marks H3K4me1, H3K4me2, and H3K27ac were used in the analysis.40
Human Kidney Biopsy Samples
Protocols involving the use of human materials and information were approved by the Institutional Review Board at the Medical College of Wisconsin or Shanghai Jiao Tong University Affiliated Sixth People’s Hospital. Records at the pathologic archive were searched to identify subjects diagnosed with hypertensive nephrosclerosis without any other type of renal injury and subjects diagnosed with minimal or no renal injury. Kidney biopsy blocks were cut into 10-μm sections. Total RNA was extracted from three sections from each subject using miRNeasy FFPE kit (Qiagen).19
Statistical Analyses
Unless described otherwise, data were analyzed using t test or multiple-group ANOVA followed by Holm–Sidak test. P<0.05 was considered significant. Data are shown as mean±SEM.
Results
miRNA Expression in the Kidneys of the Dahl SS Rat
A small RNA deep sequencing analysis was carried out in the renal outer medulla tissue from SS rats and SS.13BN26 rats that had been maintained on a diet containing 0.4% NaCl or switched at 6 weeks of age to a diet containing 4% NaCl for 7 days. The renal outer medulla was selected for the analysis because it has been shown to play a particularly important role in the development of hypertension in several models including the SS rat.14
On average, 4.7 million reads were obtained from each small RNA sequencing library, 89.6% of which could be mapped (Supplemental Table 1). Eleven known miRNAs were differentially expressed between SS and SS.13BN26 rats on the 4% NaCl diet (Figure 1A), two of which were also differentially expressed in response to the 4% NaCl diet in SS.13BN26 rats (Figure 1B). Another five miRNAs were differentially expressed in response to the 4% NaCl diet in SS rats (Figure 1C). Unadjusted P<0.05 was used as a screening criterion for identifying possible differential expression. Complete data from the rat small RNA deep sequencing analysis are available in the Supplemental Material.
Figure 1.
miRNAs differentially expressed in the renal outer medulla of SS and SS.13BN26 rats. Rats were on a 0.4% NaCl diet since weaning or switched to a 4% NaCl diet for 7 days at 6 weeks of age. Each group included nine rats that were sequenced in three pools of three rats each. miRNAs with unadjusted P<0.05 for the comparison of (A) SS versus SS.13BN26 rats on the 4% NaCl diet, (B) SS.13BN26 rats on 0.4% versus 4% NaCl diet, and (C) SS rats on the two diets are shown. (D) Real-time PCR verification of higher abundance of miR-214-3p in SS rats than SS.13BN26 rats on the 4% NaCl diet for 7 days. n=5; *P<0.05. RPM, reads per million; rRNA, ribosomal RNA.
miR-214-3p was the only differentially expressed miRNA for which the encoding gene is located in the genomic segment substituted in the congenic SS.13BN26 rat. Real-time PCR analysis of additional samples not used in the deep sequencing analysis confirmed that miR-214-3p in the renal outer medulla was significantly more abundant in SS rats on the 4% NaCl diet compared with SS.13BN26 rats (Figure 1D). miR-214-3p tended to be upregulated in SS rats on the 4% NaCl diet compared with the 0.4% NaCl diet, which just missed the significance cutoff in the miR-seq analysis (P=0.06). The passenger strand of miR-214, miR-214–5p, was not detected in the renal medulla of SS rats. For brevity, miR-214-3p was shown as miR-214 in some of the figures in this article. Other miRNA genes located in the congenic genomic segment (miR-199a, miR-3120, miR-488, and miR-1843b) were not significantly differentially expressed, and some barely detectable (Supplemental Material).
miR-214-3p abundance was significantly higher in SS rats in several other tissues including the renal cortex, liver, and heart (Supplemental Figure 1). In spontaneously hypertensive rats, renal abundance of miR-214-3p was significantly higher than in Wistar Kyoto rats (Supplemental Figure 2).
Kidney-Specific Inhibition of miR-214-3p Attenuates Salt-Induced Hypertension in SS Rats
We used a renal medullary interstitial injection approach to deliver locked nucleic acid–modified anti-miR oligonucleotides to the remaining kidney of uninephrectomized, conscious SS rats. Uninephrectomy was necessary for ensuring the kidney receiving the interstitial injection was the sole determinant of renal function of the rat. Control rats were similarly prepared and received a scrambled anti-miR. On day nine after the injection of anti–miR-214-3p, the abundance of miR-214-3p detectable with real-time PCR was significantly decreased by approximately 50% in the renal outer medulla (Figure 2A). The abundance of detectable miR-214-3p tended to decrease in the renal cortex, although the decrease did not reach statistical significance. miR-214-3p abundance in the liver was not affected (Figure 2A).
Figure 2.
Inhibition of miR-214-3p specifically in the kidney attenuates hypertension and albuminuria in SS rats. (A) miR-214-3p abundance in three kidney regions and the liver of SS rats 9 days after receiving a renal outer medullary interstitial injection of anti–miR-214 or a scrambled control anti-miR. n=5; *P<0.05 versus scrambled anti-miR. (B) Mean arterial pressure (MAP), (C) systolic BP (SBP), (D) diastolic BP (DBP), (E) heart rate (HR), and (F) albuminuria in SS rats with kidney-specific inhibition of miR-214-3p are shown. n=6 for scrambled anti-miR and n=7 for anti–miR-214 for MAP, SBP, DBP, and HR; n=5 for albuminuria; *P<0.05 versus scrambled anti-miR. rRNA, ribosomal RNA.
Significant attenuation of the increase of mean arterial pressure in rats with miR-214-3p inhibition in the kidney was detected starting on day six after the injection of anti-miR and the switch to the 4% NaCl diet (Figure 2B). The difference in mean arterial pressure reached 18 mm Hg by day nine on the 4% NaCl diet. Increases of systolic and diastolic BPs were similarly attenuated (Figure 2, C and D). Heart rate was largely not affected by the miR-214-3p inhibition (Figure 2E). The increase of 24-hour urinary excretion of albumin, prominent in SS rats as well as patients with salt-sensitive hypertension including black patients with hypertension,7,16,17 was significantly attenuated (Figure 2F).
We treated another group of SS rats for 5 days to examine whether miR-214-3p inhibition occurred before a significant effect on BP was observed. On day five, miR-214-3p in the renal outer medulla was significantly reduced to 39%±5% of the levels in rats treated with scrambled anti-miR (n=6–7; P<0.05). The detectable miR-214-3p in the renal cortex was also significantly reduced to 32%±12% of the control (P<0.05). No significant change was observed in renal inner medulla (110%±17% of control; NS).
The mRNA abundances of inflammatory marker genes TNFα, monocyte chemoattractant protein 1 (MCP1), IL-6, and IL-1 were significantly reduced on day nine, but not day five, after the anti–miR-214-3p treatment and the initiation of the 4% NaCl diet, mostly in the renal outer medulla (Supplemental Figure 3). Fibrosis-related genes α–smooth muscle actin (αSMA) and collagen IVα (Col4a) were downregulated in the renal outer medulla but not in the renal cortex on day nine and not in either kidney region on day five (Supplemental Figure 4).
eNOS Is a Direct Target of miR-214-3p
eNOS is a predicted target for miR-214-3p (Figure 3A). A 3′UTR reporter assay confirmed that miR-214-3p targeted the 3′UTR of eNOS mRNA (Figure 3B). Mutational analysis confirmed that the targeting was mediated by the binding of the seed region sequence (Figure 3, A and B). Treatment of cultured human renal epithelial cells with a mimic of miR-214-3p resulted in significant downregulation of eNOS mRNA (Figure 3C) and protein (Supplemental Figure 5) abundance. eNOS protein abundance in the renal outer medulla was significantly increased in SS rats on day five after receiving the renal interstitial injection of anti–miR-214-3p (Figure 3D). eNOS was not consistently detected in the renal cortex in any of the groups, which might be due to a variety of biologic or technical factors.
Figure 3.
miR-214-3p targets eNOS. (A) Sequence alignment of miR-214-3p, its predicted binding site in the 3′UTR of eNOS mRNA, and a point-mutated (PM) 3′UTR segment. (B) Effect of anti–miR-214-3p on the activity of luciferase with a normal or mutated segment of the eNOS 3′UTR transfected into HeLa cells. Luciferase activity was normalized by cotransfected β-gal. n=9 and n=8 for the native UTR and n=5 and n=3 for the mutated UTR; *P<0.05 versus scrambled anti-miR (control). (C) Effect of an miR-214-3p mimic on eNOS mRNA abundance in cultured human renal epithelial cells. n=3; *P<0.05. (D) Inhibition of miR-214-3p in the renal outer medulla resulted in significant upregulation of eNOS protein abundance. Western blot analysis of eNOS protein was performed on day five after kidney-specific delivery of scrambled anti-miR or anti–miR-214-3p as described in Figure 2. Western blot signals of eNOS were normalized by Coomassie blue staining of the membrane. n=5–7; *P<0.05.
We examined several rat genes involved in the renin-angiotensin system, including Ren, Agt, Ace, Ace2, Atp6ap2, Agtr1, and Agtr2. None of them is a predicted target of miR-214-3p, except a minor transcript of Ace.
The Antihypertensive Effect of Renal miR-214-3p Inhibition Is Abrogated in SS Rats with eNOS Insufficiency
One would expect the antihypertensive effect of miR-214-3p inhibition to be blunted in SS rats lacking eNOS if the effect was mediated by targeting eNOS. We developed a Nos3 mutant rat strain on the genetic background of the SS rat. Homozygous disruption of Nos3 resulted in developmental abnormalities and the rats had to be euthanized early in life for ethical reasons. SS rats with heterozygous disruption of Nos3 (SS-Nos3+/−) were used for subsequent studies.
Renal medullary interstitial administration of anti–miR-214-3p in SS-Nos3+/− rats resulted in a significant reduction of the abundance of detectable miR-214-3p particularly in the renal outer medulla (Figure 4A). In contrast to the antihypertensive effect of miR-214-3p inhibition observed in SS rats shown in Figure 2, renal inhibition of miR-214-3p did not significantly attenuate salt-induced hypertension or albuminuria in SS-Nos3+/− rats (Figure 4, B–F).
Figure 4.
Heterozygous disruption of eNOS gene abrogates the effect of renal inhibition of miR-214-3p on hypertension. (A) miR-214-3p abundance in three kidney regions and the liver of SS-Nos3+/− rats 9 days after receiving a renal medullary interstitial injection of anti–miR-214 or a scrambled control anti-miR. n=5 and n=6; *P<0.05 versus scrambled anti-miR. (B) Mean arterial pressure (MAP), (C) systolic BP (SBP), (D) diastolic BP (DBP), (E) heart rate (HR), and (F) albuminuria in SS-Nos3+/− rats with kidney-specific inhibition of miR-214-3p are shown. n=5 for scrambled anti-miR and n=8 for anti–miR-214; *P<0.05 versus scrambled anti-miR. (G) eNOS protein abundance in the renal outer medulla was significantly lower in SS-Nos3+/− rats than in wild-type SS rats even after treatment with anti–miR-214-3p. The samples from the wild-type SS and SS-Nos3+/− rats were analyzed on the same western blot gel, although not right next to each other. n=5–6; #P<0.05 versus wild-type SS rats receiving scrambled anti-miR. rRNA, ribosomal RNA.
eNOS protein abundance in the renal outer medulla of the SS-Nos3+/− rats treated with the scrambled anti-miR was approximately 50% that of wild-type SS rats treated with the scrambled anti-miR (Figure 4G), consistent with a haploinsufficiency of eNOS resulting from the heterozygous disruption of Nos3. The anti–miR-214-3p treatment resulted in a slight increase of eNOS abundance in SS-Nos3+/− rats (P=0.08). However, eNOS levels in SS-Nos3+/− rats treated with anti–miR-214-3p remained significantly lower than wild-type SS rats (Figure 4G).
miR-214-3p Is Upregulated in Kidney Biopsy Samples from Patients with Hypertension
A small RNA deep sequencing analysis was performed in sections of human kidney biopsy samples diagnosed with moderate-to-severe arterial or arteriolar nephrosclerosis consistent with hypertensive nephropathy but without any pathologic findings that would indicate the presence of any other type of kidney disease (n=8). Kidney biopsy samples without any significant pathologic findings were used as controls (n=8). Limited clinical information was available because of the historical nature of these specimens (Supplemental Table 2).
The complete dataset from the human small RNA deep sequencing analysis is available in the Supplemental Material. Nearly 10 million reads were obtained from each small RNA sequencing library, 34% of which could be mapped (Supplemental Table 3). The low mapping rates were likely because the specimens were formalin-fixed and paraffin-embedded and the quantity of extracted RNA was low. We searched for known human miRNAs that were relatively abundant (average abundance in the top 30% of all detected miRNAs) and appeared to be differentially expressed (P<0.02). Of 505 known human miRNAs detected, 22 met these criteria (Figure 5A).
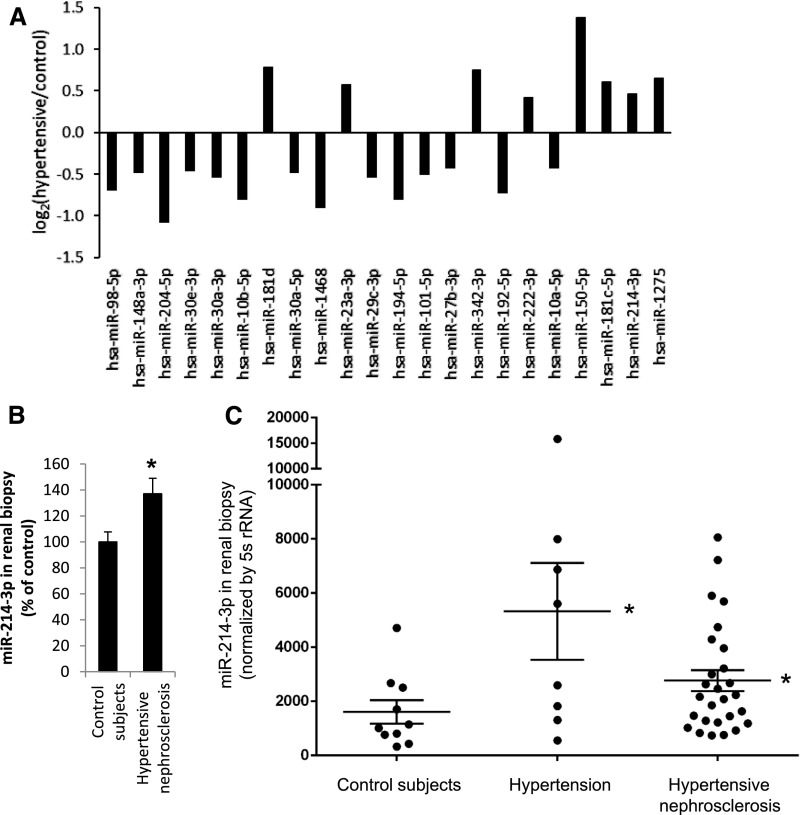
Figure 5.
MicroRNAs differentially expressed in kidney biopsy specimens from patients with hypertension. (A) miRNAs that were among the top 30% most abundant miRNAs in the kidney biopsy samples analyzed appeared to be differentially expressed in patients with hypertensive nephrosclerosis (P<0.02 versus control). n=8 subjects per group. (B) Real-time PCR verification of higher abundance of miR-214-3p in kidney biopsy samples with hypertensive nephrosclerosis. *P<0.05. (C) Higher abundance of miR-214-3p in kidney biopsy samples from a second cohort of patients with hypertension or hypertensive nephrosclerosis. miR-214-3p was measured by real-time PCR and normalized by 5s rRNA. See Supplemental Table 4 and the text for characteristics of the patients and biopsy specimens. n=10 control subjects, n=8 hypertension, and n=32 hypertensive nephrosclerosis; *P<0.05 versus control subjects. rRNA, ribosomal RNA.
Four miRNAs were identified as differentially expressed in both the human and rat studies, all of which showed changes that were directionally consistent between human and rat (see Figures 1 and 5). miR-214-3p was upregulated and miR-194–5p and miR-192–5p were downregulated in kidneys of patients with hypertensive nephrosclerosis and high-salt–fed SS rats compared with control human subjects and high-salt–fed SS.13BN26 rats, respectively. miR-222–3p was upregulated in the patients and SS rats fed the 4% salt diet compared with the 0.4% salt diet. Real-time PCR analysis confirmed that miR-214-3p was significantly upregulated in kidney biopsy samples with hypertensive nephrosclerosis (Figure 5B).
A second collection of kidney biopsy specimens was obtained for validation. Demographic and clinical characteristics of the patients in this cohort are shown in Supplemental Table 4. miR-214-3p abundance was significantly higher in kidney biopsy specimens from patients with hypertension but there were no overt renal pathologic changes (n=8) or patients diagnosed with pathologically supported hypertensive nephrosclerosis (n=32) compared with control subjects (n=10) (Figure 5C).
miR-214 Gene Is Located in Proximity to a BP-Associated SNP in the Human Genome
We performed a systematic bioinformatic analysis of human SNPs associated with BP. Of 283 sentinel SNPs that have been found to be associated with BP in GWASs, 43 were exonic and nonsynonymous, 24 exonic and synonymous, 152 intronic, and 64 intergenic including eight in promoter regions (Figure 6A, Supplemental Material). Of the 216 intronic or intergenic SNPs, 153 (71%) were located in potential enhancer regions and the LD blocks of another 49 (23%) overlapped with potential enhancer regions (Figure 6B).
Figure 6.
MicroRNA genes in genomic proximity to human BP-associated SNPs. (A) Categorization of SNPs associated with BP in humans. Two hundred eighty-three BP-associated sentinel SNPs identified in several recent GWASs were analyzed. (B) Most of the 216 BP-associated, intronic, or intergenic SNPs were in potential enhancer regions. “LD enhancer” refers to BP-associated sentinel SNPs that were not in potential enhancer regions, but one or more SNPs in LD with the sentinel SNP were. (C) Genomic relationships between coding sequences for 1870 miRNA precursors and BP-associated SNPs. (D) Several miRNAs differentially expressed in kidney biopsy specimens with hypertensive nephrosclerosis (see Figure 5A) were in genomic proximity to BP-associated SNPs. H3K4me and H3K27ac are histone marks of enhancers. (E) H3K4Me1 and H3K27Ac tracks overlaying the LD block for rs12405515 in the human genome. The LD block is in proximity to the miR-214 gene (see (D)). ENCODE, Encyclopedia of DNA Elements Project.
Analysis of the 283 SNPs and 1870 human miRNA precursors identified 16 miRNA precursors that were located within LD blocks of 16 BP-associated SNPs (Figure 6C, Supplemental Material). None of the 283 BP-associated SNPs or SNPs in LD with them were directly located within DNA segments encoding any miRNA precursor or predicted miRNA binding sites in 3′UTRs of mRNAs. Enhancers could regulate the expression of genes located up to 1 million base pairs away from them in a cis-acting manner,41 which is the genomic interval typically used in the search for expression quantitative trait loci (eQTL). eQTL data were not available for miRNAs in tissues directly relevant to BP regulation. However, 403 miRNA precursors were located within 1 million base pairs of one or more of 197 BP-associated SNPs (Figure 6C, Supplemental Material).
The genes encoding the precursors of miR-214 and four other differentially expressed miRNAs (see Figure 5A), including miR-194–2, miR-192, miR-10a, and miR-150, are located <1 million base pairs away from one or more BP-associated SNPs (Figure 6D). Moreover, the transcript that harbors pre–miR-214, DNM3os, is transcribed from a DNA strand complementary to part of DNM3, the gene that harbors the BP-associated SNP rs12405515. Another differentially expressed miRNA, miR-29c, is located in the LD block of an intergenic, BP-associated SNP (Figure 6D). These SNPs or SNPs in LD with these SNPs are in potential enhancer regions according to histone marks characteristic of enhancers (Figure 6, D and E).
Discussion
The findings of this study indicate that renal miR-214-3p functionally and potentially genetically contributes to the development of hypertension in SS rats and possibly in humans (Figure 7).
Figure 7.
Summary of approaches and findings. Findings in humans and animal models from this study suggest that miR-214-3p in the kidney functionally and potentially genetically contributes to the development of hypertension.
miR-214-3p is a highly conserved miRNA and regulates tumor progression and the development of several tissues.42–48 Much less is known about the role of miR-214 in cardiovascular and renal diseases. miR-214 may inhibit or promote endothelial angiogenesis in different settings and has been shown to be profibrotic in the kidney and antifibrotic in the heart, highlighting the tissue specificity of its effect.49–52 Our data from humans, two rat models, and multiple tissues indicate that miR-214-3p is broadly upregulated in hypertension. The kidney-targeted inhibition approach used in this study enables us to demonstrate a role of renal outer medullary miR-214-3p in contributing to salt-induced hypertension. It remains to be determined whether miR-214-3p in other tissues also functionally contributes to hypertension.
Targeting of eNOS is a plausible explanation for the hypertensive effect of renal miR-214-3p (Figure 7). The homeostatic mechanisms regulating renal regional blood flow and tubular transport are compromised in SS rats.14,53 The homeostatic impairment in SS rats might be in part caused by NO insufficiencies. NO levels are reduced in the kidneys of SS rats.54 The NO substrate L-arginine, given either systemically or locally in the renal medulla, as well as other approaches that elevate NO levels, has potent effects in attenuating hypertension in SS rats.54–56 Previous studies from our laboratory have shown broad expression of miR-214-3p in glomeruli, tubules, and vascular cells, many of which are known to express eNOS.12,13,19 Importantly, miR-214-3p directly targets eNOS, and the antihypertensive effect of miR-214-3p inhibition was lost in SS-Nos3+/− rats, suggesting that it requires an intact capacity of eNOS expression. Although the findings in SS-Nos3+/− rats might be open to alternative interpretations, the evidence taken together indicates that targeting of eNOS contributes to the hypertensive effect of miR-214-3p. The discernable role of eNOS targeting does not rule out the possibility that the hypertensive effect of renal miR-214-3p might involve other target genes.
The majority of SNPs identified by GWASs as being associated with BP or other complex traits are located in noncoding regions of the genome. Understanding how these SNPs influence cellular function, physiology, and disease is a major challenge.1,57 It has not been possible to obtain kidney or blood vessel specimens from a sufficiently large number of human subjects for a properly powered eQTL analysis. Analysis of miRNA abundance in body fluids could be done in large numbers of subjects and generate useful biomarkers, although it might be difficult to interpret the mechanistic relevance of body fluid miRNAs to hypertension as we recently discussed.58 The role of sequence variations in complex, polygenic traits such as hypertension depends highly on genomic, environmental, and cellular context.34,57,59
Despite the inevitable limitations of the human study, the two-pronged approach nominated miR-214 as one of the top-ranked miRNAs in human hypertension, which warrants further investigation. Other miRNAs nominated by the two-pronged approach include a member of the miR-29 family and miR-192, the physiologic significance of which in hypertension was supported by our previous studies13,32
Disclosures
None.
Supplementary Material
Acknowledgments
This work was supported by National Institutes of Health grants HL082798, HL121233, HL111580, HL125409, and GM066730 (to M.L.); and the National Natural Science Foundation of China (81570603 and 81770741 to F.W.).
Y.L., K.U., F.W., P.L., J.L., A.M.W., K.R.R., Y.K., H.L., J.N., and M.L. performed experiments or analyzed data. A.M.G. provided critical reagents and analytic input. M.L. designed the study. M.L. drafted the manuscript with assistance from Y.L., F.W., and A.M.W. All authors edited or approved the manuscript.
Footnotes
Published online ahead of print. Publication date available at www.jasn.org.
This article contains supplemental material online at http://jasn.asnjournals.org/lookup/suppl/doi:10.1681/ASN.2018020117/-/DCSupplemental.
References
- 1.Kotchen TA, Cowley AW Jr, Liang M: Ushering hypertension into a new era of precision medicine. JAMA 315: 343–344, 2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Padmanabhan S, Caulfield M, Dominiczak AF: Genetic and molecular aspects of hypertension. Circ Res 116: 937–959, 2015 [DOI] [PubMed] [Google Scholar]
- 3.Ehret GB, Ferreira T, Chasman DI, Jackson AU, Schmidt EM, Johnson T, et al.; CHARGE-EchoGen consortium; CHARGE-HF consortium; Wellcome Trust Case Control Consortium : The genetics of blood pressure regulation and its target organs from association studies in 342,415 individuals. Nat Genet 48: 1171–1184, 2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Liu C, Kraja AT, Smith JA, Brody JA, Franceschini N, Bis JC, et al.; CHD Exome+ Consortium; ExomeBP Consortium; GoT2DGenes Consortium; T2D-GENES Consortium; Myocardial Infarction Genetics and CARDIoGRAM Exome Consortia; CKDGen Consortium : Meta-analysis identifies common and rare variants influencing blood pressure and overlapping with metabolic trait loci. Nat Genet 48: 1162–1170, 2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Surendran P, Drenos F, Young R, Warren H, Cook JP, Manning AK, et al.; CHARGE-Heart Failure Consortium; EchoGen Consortium; METASTROKE Consortium; GIANT Consortium; EPIC-InterAct Consortium; Lifelines Cohort Study; Wellcome Trust Case Control Consortium; Understanding Society Scientific Group; EPIC-CVD Consortium; CHARGE+ Exome Chip Blood Pressure Consortium; T2D-GENES Consortium; GoT2DGenes Consortium; ExomeBP Consortium; CHD Exome+ Consortium : Trans-ancestry meta-analyses identify rare and common variants associated with blood pressure and hypertension. Nat Genet 48: 1151–1161, 2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Warren HR, Evangelou E, Cabrera CP, Gao H, Ren M, Mifsud B, et al.; International Consortium of Blood Pressure (ICBP) 1000G Analyses; BIOS Consortium; Lifelines Cohort Study; Understanding Society Scientific group; CHD Exome+ Consortium; ExomeBP Consortium; T2D-GENES Consortium; GoT2DGenes Consortium; Cohorts for Heart and Ageing Research in Genome Epidemiology (CHARGE) BP Exome Consortium; International Genomics of Blood Pressure (iGEN-BP) Consortium; UK Biobank CardioMetabolic Consortium BP working group : Genome-wide association analysis identifies novel blood pressure loci and offers biological insights into cardiovascular risk. Nat Genet 49: 403–415, 2017. 28135244 [Google Scholar]
- 7.Kotchen TA, Cowley AW Jr, Frohlich ED: Salt in health and disease--a delicate balance. N Engl J Med 368: 1229–1237, 2013 [DOI] [PubMed] [Google Scholar]
- 8.Guyton AC: Blood pressure control--special role of the kidneys and body fluids. Science 252: 1813–1816, 1991 [DOI] [PubMed] [Google Scholar]
- 9.Cowley AW Jr, Roman RJ: The role of the kidney in hypertension. JAMA 275: 1581–1589, 1996 [PubMed] [Google Scholar]
- 10.Liu Y, Taylor NE, Lu L, Usa K, Cowley AW Jr, Ferreri NR, et al.: Renal medullary microRNAs in Dahl salt-sensitive rats: miR-29b regulates several collagens and related genes. Hypertension 55: 974–982, 2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Marques FZ, Campain AE, Tomaszewski M, Zukowska-Szczechowska E, Yang YH, Charchar FJ, et al.: Gene expression profiling reveals renin mRNA overexpression in human hypertensive kidneys and a role for microRNAs. Hypertension 58: 1093–1098, 2011 [DOI] [PubMed] [Google Scholar]
- 12.Kriegel AJ, Baker MA, Liu Y, Liu P, Cowley AW Jr, Liang M: Endogenous microRNAs in human microvascular endothelial cells regulate mRNAs encoded by hypertension-related genes. Hypertension 66: 793–799, 2015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Widlansky ME, Jensen DM, Wang J, Liu Y, Geurts AM, Kriegel AJ, et al.: miR-29 contributes to normal endothelial function and can restore it in cardiometabolic disorders. EMBO Mol Med 10: e8046, 2018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Cowley AW, Jr: Renal medullary oxidative stress, pressure-natriuresis, and hypertension. Hypertension 52: 777–786, 2018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Rapp JP: Genetic analysis of inherited hypertension in the rat. Physiol Rev 80: 135–172, 2000 [DOI] [PubMed] [Google Scholar]
- 16.Lu L, Li P, Yang C, Kurth T, Misale M, Skelton M, et al.: Dynamic convergence and divergence of renal genomic and biological pathways in protection from Dahl salt-sensitive hypertension. Physiol Genomics 41: 63–70, 2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Feng D, Yang C, Geurts AM, Kurth T, Liang M, Lazar J, et al.: Increased expression of NAD(P)H oxidase subunit p67(phox) in the renal medulla contributes to excess oxidative stress and salt-sensitive hypertension. Cell Metab 15: 201–208, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Liu Y, Liu P, Yang C, Cowley AW Jr, Liang M: Base-resolution maps of 5-methylcytosine and 5-hydroxymethylcytosine in Dahl S rats: Effect of salt and genomic sequence. Hypertension 63: 827–838, 2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Baker MA, Davis SJ, Liu P, Pan X, Williams AM, Iczkowski KA, et al.: Tissue-specific microRNA expression patterns in four types of kidney disease. J Am Soc Nephrol 28: 2985–2992, 2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kriegel AJ, Liu Y, Liu P, Baker MA, Hodges MR, Hua X, et al.: Characteristics of microRNAs enriched in specific cell types and primary tissue types in solid organs. Physiol Genomics 45: 1144–1156, 2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tian Z, Greene AS, Pietrusz JL, Matus IR, Liang M: MicroRNA-target pairs in the rat kidney identified by microRNA microarray, proteomic, and bioinformatic analysis. Genome Res 18: 404–411, 2008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Liu Y, Singh RJ, Usa K, Netzel BC, Liang M: Renal medullary 11 beta-hydroxysteroid dehydrogenase type 1 in Dahl salt-sensitive hypertension. Physiol Genomics 36: 52–58, 2008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Liu Y, Mladinov D, Pietrusz JL, Usa K, Liang M: Glucocorticoid response elements and 11 beta-hydroxysteroid dehydrogenases in the regulation of endothelial nitric oxide synthase expression. Cardiovasc Res 81: 140–147, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Tian Z, Liu Y, Usa K, Mladinov D, Fang Y, Ding X, et al.: Novel role of fumarate metabolism in dahl-salt sensitive hypertension. Hypertension 54: 255–260, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Taylor NE, Glocka P, Liang M, Cowley AW Jr: NADPH oxidase in the renal medulla causes oxidative stress and contributes to salt-sensitive hypertension in Dahl S rats. Hypertension 47: 692–698, 2006 [DOI] [PubMed] [Google Scholar]
- 26.Mattson DL, Lu S, Nakanishi K, Papanek PE, Cowley AW Jr: Effect of chronic renal medullary nitric oxide inhibition on blood pressure. Am J Physiol 266: H1918–H1926, 1994 [DOI] [PubMed] [Google Scholar]
- 27.Kassab S, Novak J, Miller T, Kirchner K, Granger J: Role of endothelin in mediating the attenuated renal hemodynamics in Dahl salt-sensitive hypertension. Hypertension 30: 682–686, 1997 [DOI] [PubMed] [Google Scholar]
- 28.Gross JM, Berndt TJ, Knox FG: Effect of serotonin receptor antagonist on phosphate excretion. J Am Soc Nephrol 11: 1002–1007, 2000 [DOI] [PubMed] [Google Scholar]
- 29.Padia SH, Kemp BA, Howell NL, Siragy HM, Fournie-Zaluski MC, Roques BP, et al.: Intrarenal aminopeptidase N inhibition augments natriuretic responses to angiotensin III in angiotensin type 1 receptor-blocked rats. Hypertension 49: 625–630, 2007 [DOI] [PubMed] [Google Scholar]
- 30.Li N, Chen L, Yi F, Xia M, Li PL: Salt-sensitive hypertension induced by decoy of transcription factor hypoxia-inducible factor-1alpha in the renal medulla. Circ Res 102: 1101–1108, 2008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kriegel AJ, Liu Y, Cohen B, Usa K, Liu Y, Liang M: MiR-382 targeting of kallikrein 5 contributes to renal inner medullary interstitial fibrosis. Physiol Genomics 44: 259–267, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Mladinov D, Liu Y, Mattson DL, Liang M: MicroRNAs contribute to the maintenance of cell-type-specific physiological characteristics: miR-192 targets Na+/K+-ATPase β1. Nucleic Acids Res 41: 1273–1283, 2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kriegel AJ, Fang Y, Liu Y, Tian Z, Mladinov D, Matus IR, et al.: MicroRNA-target pairs in human renal epithelial cells treated with transforming growth factor beta 1: A novel role of miR-382. Nucleic Acids Res 38: 8338–8347, 2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Usa K, Liu Y, Geurts AM, Cheng Y, Lazar J, Baker MA, et al.: Elevation of fumarase attenuates hypertension and can result from a nonsynonymous sequence variation or increased expression depending on rat strain. Physiol Genomics 49: 496–504, 2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Geurts AM, Cost GJ, Freyvert Y, Zeitler B, Miller JC, Choi VM, et al.: Knockout rats via embryo microinjection of zinc-finger nucleases. Science 325: 433, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wang F, Zhang G, Lu Z, Geurts AM, Usa K, Jacob HJ, et al.: Antithrombin III/SerpinC1 insufficiency exacerbates renal ischemia/reperfusion injury. Kidney Int 88: 796–803, 2015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lawrence M, Huber W, Pagès H, Aboyoun P, Carlson M, Gentleman R, et al.: Software for computing and annotating genomic ranges. PLOS Comput Biol 9: e1003118, 2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ward LD, Kellis M: HaploReg: A resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res 40: D930–D934, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kozomara A, Griffiths-Jones S: miRBase: Annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res 42: D68–D73, 2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Heinz S, Romanoski CE, Benner C, Glass CK: The selection and function of cell type-specific enhancers. Nat Rev Mol Cell Biol 16: 144–154, 2015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Pennacchio LA, Bickmore W, Dean A, Nobrega MA, Bejerano G: Enhancers: Five essential questions. Nat Rev Genet 14: 288–295, 2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Flynt AS, Li N, Thatcher EJ, Solnica-Krezel L, Patton JG: Zebrafish miR-214 modulates Hedgehog signaling to specify muscle cell fate. Nat Genet 39: 259–263, 2007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Juan AH, Kumar RM, Marx JG, Young RA, Sartorelli V: Mir-214-dependent regulation of the polycomb protein Ezh2 in skeletal muscle and embryonic stem cells. Mol Cell 36: 61–74, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Penna E, Orso F, Cimino D, Tenaglia E, Lembo A, Quaglino E, et al.: microRNA-214 contributes to melanoma tumour progression through suppression of TFAP2C. EMBO J 30: 1990–2007, 2011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Sehic A, Risnes S, Khuu C, Khan QE, Osmundsen H: Effects of in vivo transfection with anti-miR-214 on gene expression in murine molar tooth germ. Physiol Genomics 43: 488–498, 2011 [DOI] [PubMed] [Google Scholar]
- 46.Ahmed MI, Alam M, Emelianov VU, Poterlowicz K, Patel A, Sharov AA, et al.: MicroRNA-214 controls skin and hair follicle development by modulating the activity of the Wnt pathway. J Cell Biol 207: 549–567, 2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Nakagawa N, Xin C, Roach AM, Naiman N, Shankland SJ, Ligresti G, et al.: Dicer1 activity in the stromal compartment regulates nephron differentiation and vascular patterning during mammalian kidney organogenesis. Kidney Int 87: 1125–1140, 2015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Li D, Liu J, Guo B, Liang C, Dang L, Lu C, et al.: Osteoclast-derived exosomal miR-214-3p inhibits osteoblastic bone formation. Nat Commun 7: 10872, 2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.van Mil A, Grundmann S, Goumans MJ, Lei Z, Oerlemans MI, Jaksani S, et al.: MicroRNA-214 inhibits angiogenesis by targeting Quaking and reducing angiogenic growth factor release. Cardiovasc Res 93: 655–665, 2012 [DOI] [PubMed] [Google Scholar]
- 50.van Balkom BW, de Jong OG, Smits M, Brummelman J, den Ouden K, de Bree PM, et al.: Endothelial cells require miR-214 to secrete exosomes that suppress senescence and induce angiogenesis in human and mouse endothelial cells. Blood 121: 3997–4006, S1–S15, 2013 [DOI] [PubMed] [Google Scholar]
- 51.Denby L, Ramdas V, Lu R, Conway BR, Grant JS, Dickinson B, et al.: MicroRNA-214 antagonism protects against renal fibrosis. J Am Soc Nephrol 25: 65–80, 2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Aurora AB, Mahmoud AI, Luo X, Johnson BA, van Rooij E, Matsuzaki S, et al.: MicroRNA-214 protects the mouse heart from ischemic injury by controlling Ca2+ overload and cell death. J Clin Invest 122: 1222–1232, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Roman RJ: Abnormal renal hemodynamics and pressure-natriuresis relationship in Dahl salt-sensitive rats. Am J Physiol 251: F57–F65, 1986 [DOI] [PubMed] [Google Scholar]
- 54.Hou E, Sun N, Zhang F, Zhao C, Usa K, Liang M, et al.: Malate and aspartate increase L-arginine and nitric oxide and attenuate hypertension. Cell Reports 19: 1631–1639, 2017 [DOI] [PubMed] [Google Scholar]
- 55.Chen PY, Sanders PW: L-arginine abrogates salt-sensitive hypertension in Dahl/Rapp rats. J Clin Invest 88: 1559–1567, 1991 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Miyata N, Zou AP, Mattson DL, Cowley AW Jr: Renal medullary interstitial infusion of L-arginine prevents hypertension in Dahl salt-sensitive rats. Am J Physiol 275: R1667–R1673, 1998 [DOI] [PubMed] [Google Scholar]
- 57.Mattson DL, Liang M: Hypertension: From GWAS to functional genomics-based precision medicine. Nat Rev Nephrol 13: 195–196, 2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Touyz RM, Montezano AC, Rios F, Widlansky ME, Liang M: Redox stress defines the small artery vasculopathy of hypertension: How do we bridge the bench-to-bedside gap? Circ Res 120: 1721–1723, 2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Liang M, Cowley AW Jr, Mattson DL, Kotchen TA, Liu Y: Epigenomics of hypertension. Semin Nephrol 33: 392–399, 2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.