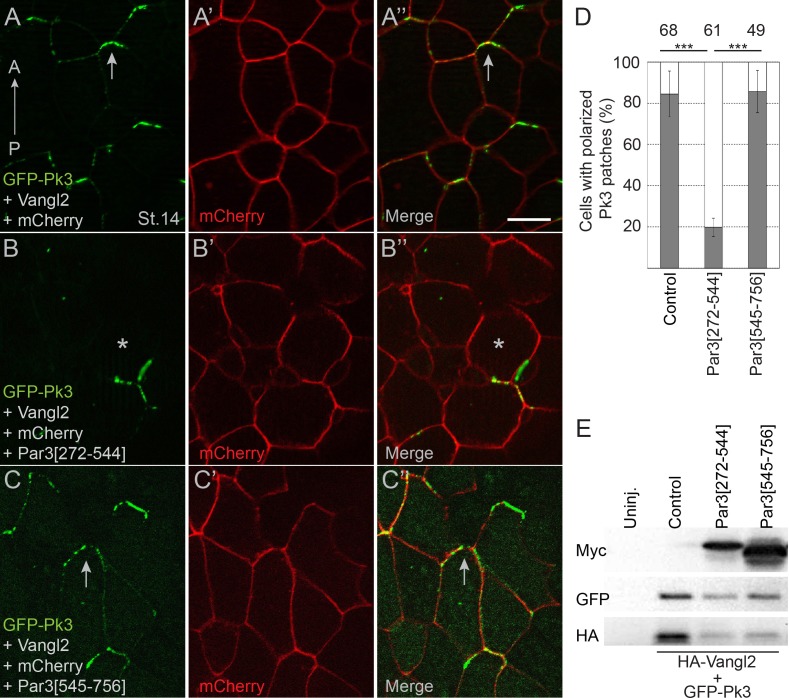
Figure 5. Pk3-interacting fragment of Par3 interferes with neural plate PCP.
Two dorsal blastomeres of 16 cell embryos were injected with RNAs encoding GFP-Pk3 (100 pg), HA-Vangl2 (25 pg) and mCherry (70 pg) without (A–A”) or with Par3[272-544] (0.5 ng) (B–B’’) or Par3[545-756] (0.5 ng) (C–C’’). Cells from embryos at stage 14 (St.14) with anteriorly polarized (arrows) and mislocalized (asterisks) GFP-Pk3 patches are shown. Anteroposterior (AP) axis of the neural plate is indicated. Scale bar, 20 µm. (D) Quantification of data in (A–C) shown as mean frequencies ± s. d. of polarized GFP-Pk3 patches in neuroepithelial cells. Total numbers of scored cells are shown above each bar; 5 to 25 cells were scored per embryo with five embryos taken for each experimental condition, statistical significance was determined by two-tailed Student’s t-test, p<0.001. Data are representative of two experiments. (E) Protein expression levels were assessed in stage 14 embryos by immunoblotting with anti-Myc, anti-GFP and anti-HA antibodies. Control, embryos injected with HA-Vangl2 and GFP-Pk3 RNAs without Par3 constructs, Uninj., uninjected embryos.