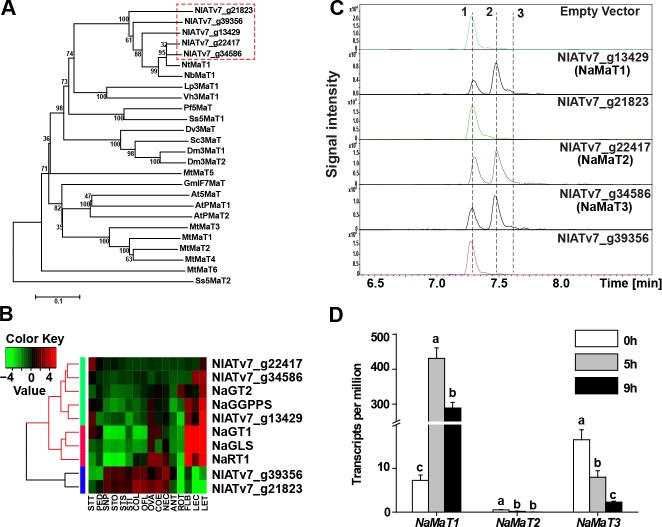
Figure 2. NaMaT1 mediates DTG malonylation in N. attenuata.
(A) Phylogenetic analysis of potential N. attenuata malonyltransferases (genes in red dotted box) and functionally characterized malonyltransferases in other species by amino acid sequence with accession number shown in Supplementary file 1. (B) Heatmap representing the expression of malonyltransferases and reported DTG biosynthetic genes in N. attenuata. LET, leaf treated (25 hr after wounding and elicitation with M. sexta oral secretions); LEC, leaf control; STT, stem treated; PED, pedicels; SNP, style without pollination; STO, style outcrossed; STS, style selfed; STI, stigma; COL, corolla late; OFL, opening flower; OVA, ovary; COE, corolla early; NEC, nectaries; ANT, anthers; ROT, root OS-elicited; FLB, flower bud. (C) Extracted ion chromatograms of m/z 271.2420, corresponding to the DTG aglycone, of in vitro assay products of recombinant malonyltransferases. Peaks 1, 2 and 3 were identified as Lyciumoside IV, Nicotianoside I and Nicotianoside II, respectively. (D) NaMaT1, NaMaT2 and NaMaT3 transcript counts (mean + SE; n = 3) were analyzed from RNAseq data of M. sexta-attacked leaves at indicated time points. Different letters indicate significant differences among treated time points (p<0.05, one ANOVA followed by Tukey’s HSD post-hoc tests).
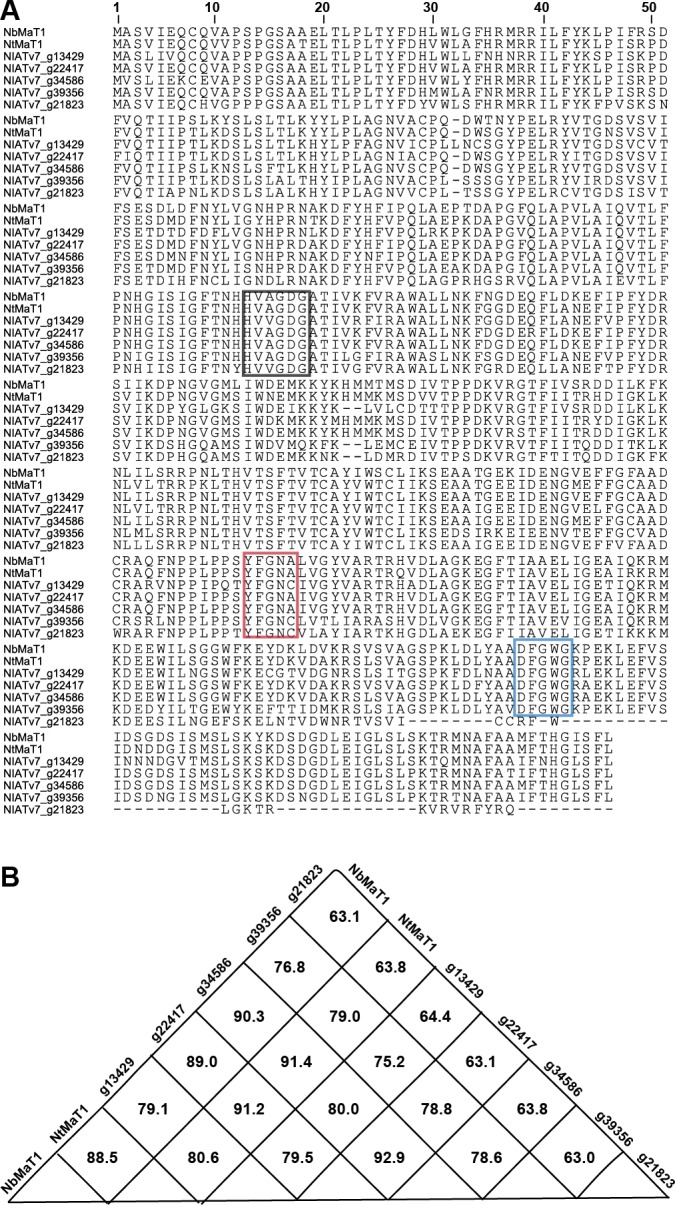
Figure 2—figure supplement 1. Analyzing protein sequences of NbMaT1, NtMaT1 and malonyltransferases in N. attenuata.