Abstract
Monoubiquitination of histone H2B (to H2Bub1) is required for downstream events including histone H3 methylation, transcription, and mRNA export. The mechanisms and players regulating these events have not yet been completely delineated. Here, we show that the conserved Ran‐binding protein Mog1 is required to sustain normal levels of H2Bub1 and H3K4me3 in Saccharomyces cerevisiae. Mog1 is needed for gene body recruitment of Rad6, Bre1, and Rtf1 that are involved in H2B ubiquitination and genetically interacts with these factors. We provide evidence that the absence of MOG1 impacts on cellular processes such as transcription, DNA replication, and mRNA export, which are linked to H2Bub1. Importantly, the mRNA export defect in mog1Δ strains is exacerbated by the absence of factors that decrease H2Bub1 levels. Consistent with a role in sustaining H2Bub and H3K4me3 levels, Mog1 co‐precipitates with components that participate in these modifications such as Bre1, Rtf1, and the COMPASS‐associated factors Shg1 and Sdc1. These results reveal a novel role for Mog1 in H2B ubiquitination, transcription, and mRNA biogenesis.
Keywords: epigenetics, H2B ubiquitination, mRNA export, transcription, yeast
Subject Categories: Chromatin, Epigenetics, Genomics & Functional Genomics; Transcription
Introduction
Eukaryotic gene expression is a complex process that is regulated at multiple levels, including nuclear and cytoplasmic events. Prior to mRNA export from the nucleus, a sophisticated crosstalk between different histone modifications acts to control transcription and its coupling to mRNA processing and export 1, 2, 3. A crucial histone modification implicated in this crosstalk is monoubiquitination of histone H2B on lysine 123 in yeast (K123) or on lysine 120 in mammals (K120) to form H2Bub1 4. H2Bub1 is involved in a variety of biological processes ranging from cellular growth to transcription initiation and elongation, and gene silencing 5. Dysregulation of this modification in higher eukaryotes leads to important pathological events such as tumorigenesis and deficiencies in cellular differentiation and development 6, 7. The E2 ubiquitin‐conjugating enzyme Rad6 catalyzes H2B monoubiquitination in conjunction with the E3 ubiquitin ligase Bre1 and the transcriptional regulatory protein Lge1 8, 9, 10, 11. H2Bub1 is present throughout the entire transcriptional units of active genes due to the association of Rad6 and Bre1 with chromatin through their interaction with activators and with RNA polymerase II (RNAPII) 12, 13. H2B is, therefore, co‐transcriptionally ubiquitinated, and its H2Bub1 levels correlate with RNAPII elongation rates 14. The monoubiquitination of H2B also requires the presence of the PAF complex (PAFc). Recent studies have shown that the Rtf1 subunit of the PAFc acts as a cofactor for H2B monoubiquitination by promoting the ability of Bre1 to stimulate Rad6 activity 15. Mutations affecting H2Bub1 levels specifically abolish di‐ and trimethylation of H3K4 and H3K79 while only marginally affecting monomethylation 16, 17. Rad6/Bre1‐dependent H2Bub1 also promotes the ubiquitination of Swd2 resulting in control of the recruitment to chromatin of Spp1, a COMPASS subunit that stimulates the overall rate of methylation of H3K4 18, 19. It is consequently thought that the PAFc affects H3K4 and H3K79 methylation through its effect on H2B ubiquitination 8, 20.
Both the addition and the removal of the ubiquitin moiety to and from H2B are required for optimal gene transcription. H2Bub1 is deubiquitinated by two ubiquitin‐specific proteases in S. cerevisiae: Ubp8 and Ubp10. Ubp8 is part of the deubiquitination module (DUBm) of the SAGA complex, which is composed of four proteins that are conserved from yeast to humans: yUbp8/hUSP22, ySus1/hENY2, ySgf73/hATXN7, and ySgf11/hATXN7L3 21, 22, 23, 24, 25. The presence of these four subunits, together with their correct assembly, is necessary for H2Bub1 deubiquitination 26, 27, 28, 29, 30, 31. Several studies have provided important insights into the crystal structure of the DUBm and have shown that the four components are remarkably intertwined 28, 31, 32. Notably, recent work in vitro has shown that the DUBm can contact the nucleosome core particle acidic patch via Sgf11 to which it competes with Rad6‐Bre1 for binding. This competition might serve to fine‐tune cellular levels of H2Bub1 genome‐wide 33.
H2Bub1 is also functionally connected with mRNA export. For instance, Sus1, a component of both SAGA DUBm and TREX‐2 complexes, plays a key role in coupling transcription activation, H2Bub1 deubiquitination, and mRNA export 34. It has also been shown that deletion of either DUBm subunits Sgf11 or Sgf73 exacerbates mRNA export defects 35, 36, while the TREX‐2 subunit Sem1 is required for SAGA‐dependent deubiquitinating activity 37. These data highlight the bidirectional functional connection between the two complexes. In addition, H2Bub1 controls the formation of export‐competent mRNPs by affecting recruitment of the different Mex67 adaptors to these complexes and by regulating the levels of the COMPASS subunit Swd2 2.
All of these findings indicate strong crosstalk between H2Bub1 and mRNP biogenesis. Here, we identified a novel component required for proper H2B monoubiquitination in yeast: Mog1, which is a conserved Ran GTPase‐binding protein that is required for nuclear protein import 38, 39, 40. We demonstrate that deletion of MOG1 affects the global levels of both H2Bub1 and H3K4me3 and the chromatin recruitment of factors implicated in their modifications. Using Genomic Run‐On (GRO), Mog1‐TAP purification, and ChIP techniques, we determined that cells lacking MOG1 showed a global impairment of transcription that is consistent with the co‐purification of Mog1 with factors involved in transcription and with Mog1 association with chromatin. Notably, defects in mRNA export are exacerbated by the combination of MOG1 deletion and mutations in H2B ubiquitination factors, suggesting that upstream processes that are partly coordinated by Mog1 affect mRNA export. This new function of Mog1, which is independent of Mog1 binding to Ran‐GTP, extends its functions beyond its well‐known role in protein import.
Results
Mog1 stimulates the monoubiquitination of H2B
Sus1 plays a critical role in coordinating histone H2B monoubiquitination, transcription, and mRNA export 35, 36. We hypothesized that identification of factors that genetically interact with SUS1 might lead to the discovery of new factors involved in this coordination. We found that mog1Δ displayed the strongest genetic interaction with SUS1 deletion in genome‐wide screenings using synthetic genetic arrays (SGA) (mog1Δ: SGA score −0.67 41, 42. Mog1 is a nuclear protein that interacts with Ran‐GTP. Mog1 stimulates the release of GTP from Ran, thereby conferring directionality to the nuclear import and export pathways 39, 40. In order to validate this genetic interaction, we created a double‐deletion mutant and analyzed the effect of MOG1 deletion on the growth of cells lacking SUS1. mog1Δ showed a negative genetic interaction with sus1Δ (Fig 1A). Next, we analyzed Gene Ontology (GO) of all genes that showed a genetic interaction with mog1Δ (Dataset EV1 ). Notably, the four members of the SAGA deubiquitination module (Sus1, Sgf73, Ubp8, and Sgf11) were the genes whose GO annotation had the highest enrichment (Table EV1), suggesting that the DUBm components are genetically linked to MOG1. To confirm these genetic interactions, double mutants were created in which mog1Δ was combined with mutants of DUBm components. Genetic interactions were evident when MOG1 deletion was combined with deletion of UBP8 or of SGF73, with the strongest effect on growth observed for the double mutant mog1Δubp8Δ (Fig 1A).
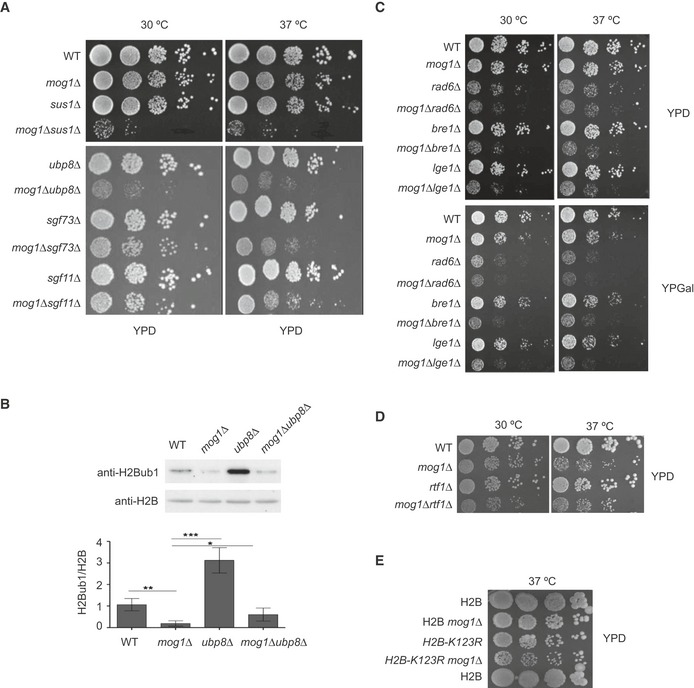
Figure 1. Monoubiquitination of H2B requires Mog1.
-
AMOG1 interacts genetically with the H2B deubiquitination machinery. Tenfold serial dilutions of the indicated strains were spotted on YPD and incubated for 2 days at the indicated temperatures.
-
BExpression levels of total H2B and H2Bub1 were analyzed in wild‐type (WT), mog1Δ, ubp8Δ, and ubp8Δmog1Δ whole‐cell lysates by Western blotting using anti‐total H2B and anti‐H2Bub1 antibodies. Bar graphs of H2Bub1 levels after H2B normalization show the mean and standard deviations of at least three independent experiments. Error bars represent SD. The P‐value was calculated using Student's t‐test (*P = 0.01–0.05; **P = 0.001–0.01; ***P < 0.001).
-
C–EThe indicated strains were diluted as described in (A) and were then spotted on YPD or YPGal and incubated for 2 days.
Since genetic interactions might potentially be indicative of functional relationships, we decided to analyze the global levels of histone H2Bub1 in a mog1Δ mutant. The absence of MOG1 significantly reduced the total level of H2Bub1as determined by Western blotting (Fig 1B). To determine whether the low levels of H2Bub1 were a consequence of low ubiquitination or of high Ubp8‐dependent deubiquitinating activity, the double mutant mog1Δubp8Δ was also analyzed. Fig 1B shows that the reduction in H2Bub1 levels observed in the absence of MOG1 was not fully rescued by UBP8 deletion, suggesting that the low H2Bub1 level in the mog1Δ strain was not UBP8‐dependent. In accordance with this result, MOG1 deletion did not affect the interactions of TAP‐tagged Ubp8 with other cellular proteins or its chromatin association (Fig EV1A and B and Table EV2). In summary, these data suggested that Mog1 affected H2B monoubiquitination rather than the opposing H2B deubiquitinating activity.
Figure EV1. Ubp8 phenotypes are not affected by MOG1 deletion.

- A representative Coomassie‐stained SDS–PAGE gel showing the pattern of proteins co‐purifying with Ubp8‐TAP from wild‐type (WT) or mog1Δ cells. Protein markers (M) of the indicated molecular weight (kDa) are shown in lane M. The list of proteins detected using mass spectrometric analysis is shown in Table EV2.
- ChIP analysis of the presence of TAP‐tagged Ubp8 at ADH1, PMA1, and YEF3 promoter (P) and coding regions (5'ORF) in WT or mog1Δ strains. The bar graphs indicate the mean and standard deviation for at least three independent experiments. Significance of the differences was calculated using Student's t‐test.
Mog1 interacts genetically with components involved in H2B monoubiquitination
To explore the interaction between Mog1 and the H2B monoubiquitination machinery, genetic assays were conducted in which MOG1 deletion was combined with individual mutants of RAD6 (E2), BRE1 (E3), or its associated protein LGE1. Growth assays indicated that mutation of MOG1 led to a synthetic sickness phenotype when combined with a deletion of any of the three genes (Fig 1C). Since H2Bub1 levels are also regulated by the PAFc 43, we combined MOG1 deletion with deletion of the PAFc subunit RTF1 and found that the resulting double mutant displayed an additive growth phenotype (Fig 1D). These results indicated that Mog1 and Rad6, Bre1, or Rtf1 act in independent pathways. However, since the absence of any of the factors Rad6, Bre1, or Rtf1 alone results in undetectable levels of H2Bub1 15, 33, it is unlikely that the growth phenotypes of the double mutants are a consequence of further reduction in H2Bub1 levels that is caused by mog1Δ. To examine whether a process in addition to H2B monoubiquitination is affected by the loss of Mog1, we analyzed the genetic interaction between mog1Δ and a histone H2B mutant (H2B‐K123R) that is specifically defective in H2B monoubiquitination 44. As shown in Fig 1E, deletion of MOG1 in the H2B‐K123R mutant resulted in a subtle growth retardation suggesting that Mog1 may affect another process (or processes) in addition to H2B monoubiquitination.
Since Mog1 participates in nuclear protein import, we asked whether partial disruption of this pathway might be responsible for the observed genetic interactions and the decreased levels of H2Bub1. To examine this possibility, Bre1 and Rad6 were GFP‐tagged and their cellular localization in mog1Δ compared to wild‐type cells was monitored using confocal microscopy. Similar to wild‐type, both Rad6 and Bre1 showed a nuclear localization in cells lacking MOG1 (Fig EV2A and B). In addition, GFP‐tagged Rtf1 and Ubp8 showed a nuclear localization in cells lacking MOG1 (Fig EV2C and D). In summary, these genetic and functional data suggested that Mog1 affects another pathway(s) in addition to H2B monoubiquitination.
Figure EV2. Mog1 is not required for Rad6, Bre1, Rtf1, or Ubp8 nuclear localization.

-
A–DRepresentative images showing the GFP signal for Rad6‐GFP (A), Bre1‐GFP (B), Rtf1‐GFP (C), and Ubp8‐GFP (D) in WT or mog1Δ strains. DAPI (DNA) and BF (bright field) pictures of the same images are shown. In all images, the scale bar corresponds to 5 μm.
Global levels of H3K4me3 decrease in the absence of MOG1
H2Bub1 promotes the presence of H3K4me3 at actively transcribing genes 45. The molecular mechanisms behind this crosstalk are complex and are postulated to require multiple factors. H3K4 is methylated by chromatin binding of Set1 and its associated factors (the COMPASS complex), which correlates with productive transcription elongation and mRNA biogenesis 45, 46. In S. cerevisiae, Set1 associates with chromatin even in the absence of H2Bub1; therefore, it was proposed that Set1 chromatin recruitment is not the primary function of H2Bub1 17, 47. However, in S. pombe, H2Bub1 directly enhances the activity of intact Set1C by stabilizing its interaction with chromatin. To determine how H3K4me3 might be affected in the absence of MOG1, differences in global trimethylation levels of H3K4 were analyzed in the presence and absence of MOG1. Western analysis revealed that levels of H3K4me3 were significantly reduced in mog1Δ whole‐cell protein extracts compared to wild‐type extracts (Fig 2A). Concomitantly, the deletion of MOG1 led to a slight decrease in trimethylation of H3K4 at promoter and 5'ORF regions of ADH1, PMA1, and YEF3 genes as assessed by ChIP analysis (Fig 2B).
Figure 2. Mog1 affects trimethylation of H3K4.

- The expression levels of H3K4me3 and H3 in total extracts of wild‐type (WT), mog1Δ, set1Δ, and spp1Δ strains were analyzed by Western blotting using the indicated antibodies. Bar graphs of H3K4me3 levels after H3 normalization show the mean and standard deviations of at least three independent experiments. The P‐value was calculated using Student's t‐test (*P = 0.01–0.05; ***P < 0.001).
- ChIP analysis of H3K4me3 presence at ADH1, PMA1, and YEF3 promoter (P) and 5'ORF in wild‐type (WT) or mog1Δ strains relative to total H3. The bar graphs indicate the mean and standard deviation for at least three independent experiments. The significance of the differences was calculated using Student's t‐test (*P = 0.01–0.05; **P = 0.001–0.01; ***P < 0.001).
- MOG1 interacts genetically with the H3 methylases Set1 and Set2. Tenfold serial dilutions of the indicated strains were spotted on YPD and incubated for 2 days at the indicated temperatures.
- Histone H3K4A and H3K4A mog1Δ strains were compared with their isogenic strains as in (C).
H2Bub1 not only promotes Set1‐dependent histone H3K4 methylation, but also Dot1‐dependent H3K79 methylation 48, and it facilitates the Set2‐mediated methylation of H3K36 on some intron‐containing genes 3. However, global levels of H3K79me3 and of H3K36me3 were not affected by the loss of MOG1 (Fig EV3A and B). Additionally, no effect on H3K4me2 levels was observed by deletion of MOG1 (Fig EV3 C). These observations are similar to previous findings that H2B mutations that specifically affect its C‐terminal helix (T122R, T122D, and R119A) had no effect on H3K4me2 or on H3K79me2 or me3 levels but did affect H3K4me3 levels 49.
Figure EV3. Mog1 is not required for H3K79me3, H3K36me3, and H3K4me2 methylations.

- The expression levels of H3K79me3 and of total H3 in total extracts from WT, mog1Δ, and bre1Δ cells were analyzed by Western blotting using the indicated antibodies.
- As in (A) except that set2Δ, WT, and mog1Δ strains were analyzed using an anti‐H3K36me3 antibody.
- As in (A) except that WT, mog1Δ, and rad1Δ strains were analyzed using an anti‐H3K4me2 antibody.
Next, we tested the potential genetic interaction between Mog1 and the methylases responsible for these modifications. A slow growth phenotype was observed when mog1Δ was combined with set1Δ or set2Δ but not when it was combined with dot1Δ (Fig 2C). Since a pronounced phenotype of mog1Δ cells was a reduction in H3K4me3 levels, we addressed whether the slow growth observed for mog1Δset1Δ was directly related to this specific lysine modification of H3. To this end, we deleted MOG1 in a H3K4A mutant background and observed that the resulting strain was synthetic sick at 33°C (Fig 2D). In summary, these genetic results suggested that the synthetic growth effects observed for mog1Δ in combination with mutants affecting H2B ubiquitination and H3 methylation cannot only be explained by the loss of these modifications. These findings are consistent with the notion that MOG1 deletion disrupts independent pathways that converge on an essential process (or processes) in yeast cells.
Mog1 absence affects H2B monoubiquitination‐dependent processes such as transcription
The above data show that Mog1 was required to sustain correct levels of H2Bub1. H2Bub1 has been implicated in various fundamental cellular functions including the regulation of transcription 50, nucleosome dynamics, and cell size control 51. Among these functions, H2Bub1 promotes the stability and advancement of replication forks by regulating nucleosome stability on newly replicated DNA 52. It has been shown that the non‐ubiquitinatable H2B mutant (K123R) and bre1Δ cells are sensitive to hydroxyurea (HU), a replication stress‐causing agent 52. To examine whether MOG1 deletion lead toward sensitivity to HU, mog1Δ or bre1Δmog1Δ cells were grown in media containing different concentrations of HU. Notably, mog1Δ cells were sensitive to hydroxyurea, and this sensitivity increased in the double mutant bre1Δmog1Δ (Fig 3A).
Figure 3. MOG1 deletion affects sensitivity to hydroxyurea and RNAPII‐dependent transcription.

-
ATenfold serial dilutions of the indicated strains were spotted on YPD containing the indicated concentrations of hydroxyurea (HU) and were incubated for 2–3 days.
-
B, C(B) mRNA synthesis rates (SR) as determined in Genomic Run‐On analysis and (C) mRNA abundance (RA), in mog1Δ compared with wild‐type (WT) cells. Bar graphs showing the average level and standard deviation of three experiments (n = 3) for SR and RA values obtained as the median of the whole gene dataset values in arbitrary units (×107) from image analysis quantification. Error bars represent SD. Significance of the differences was obtained using Student's t‐test, and statistical difference is considered as *P = 0.01–0.05. The scatter plots (right panels) show the variation of the transcription (B) or mRNA (C) levels of individual genes. Pearson correlation values are −0.039 (SR) and −0.042 (RA). This low correlation and the flat shape of the clouds on scatter plots (see red tendency line) indicates that there is no bias of the transcriptional effect with regard to expression level.
-
DBox plots of SR and RA median levels of mog1Δ cells compared with WT for all genes (Total), SAGA and TFIID‐dominated genes. These data were obtained from three independent replicates (n = 3) of the RA and SR data that were averaged as explained in the main text. The line within the box and the diplayed number represent the median of the whole data set (Total) for 351 (for RA) or 316 (SR) SAGA‐dominated genes and for the 3,903 (RA) or 3,403 (SR) TFIID‐dominated genes. The box represents the second quartile and the whiskers the first and third quartiles. Significance of the differences was calculated using Student's t‐test. No significant difference between the different groups was found. The similar variations in SRs and RAs indicate that there are no changes in mRNA half‐lives upon MOG1 deletion (shown in Fig EV4). The total number of genes with confident value in the analyses was 3,953 for RA and 3,989 for SR.
Since H2B ubiquitination is a major regulator of transcription 53, we considered whether the absence of MOG1 might interfere with global mRNA transcription. To date, no functional interactions between Mog1 and the transcriptional machinery have been reported. To address such possible interactions, we conducted Genomic Run‐On (GRO) experiments 54 to calculate the mRNA synthesis rate (SR, see Materials and Methods) for each transcript in wild‐type and mog1Δ cells. In addition to the SR, the mRNA abundance (RA) for each gene was also calculated. These results showed that the global levels of both the SR and RA were significantly lower for mog1Δ compared to wild‐type (0.746 and 0.707, respectively, see Fig 3B and C). The quantitatively similar behavior of SR and RA indicates that global mRNA stability was not appreciably different between the two yeast strains (Fig EV4). Moreover, supportive of a general role for Mog1 in affecting the transcriptional rate and mRNA abundance, similar effects were observed for transcription of SAGA‐ and TFIID‐dominated genes (Fig 3D), in agreement with the contribution of both SAGA and TFIID to the expression of nearly all yeast genes 55, 56, 57, 58. These findings led us to the conclusion that Mog1 has a genome‐wide impact on RNAPII‐transcribed mRNAs.
Figure EV4. Global mRNA stability did not change appreciably in the absence of Mog1.

Box plot showing the relative stability for mRNAs (RS) calculated for 3,450 genes by dividing the mRNA amount data (RA) by the synthesis rate data (SR) used in Fig 3. These data were obtained from three independent replicates (n = 3) of the RA and SR data that were averaged as explained in the main text. The line within the box and the displayed number represent the median of the whole data set (Total) and for 316 SAGA‐dominated genes and 3,403 TFIID‐dominated genes are shown. The box represents the second quartile and the whiskers the first and third quartiles. Significance of the differences was calculated using Student's t‐test. No significant difference between them was found.
Mog1 affects the occupancy of H2B ubiquitination‐promoting factors on transcribed genes
Ubiquitination and deubiquitination of H2B during transcription are complex processes that require the participation of multiple factors 51. Recent reports have shown that one such factor, Rtf1 (a PAFc subunit), acts as a cofactor in H2B ubiquitination by promoting the ability of Bre1 to direct Rad6 to lysines on H2B or by stimulating Rad6 catalytic activity 15. To gain insights into a possible upstream role for Mog1 in H2B ubiquitination, chromatin recruitment of Rad6, Bre1, and Rtf1 to the 5′ and 3′ coding regions of ADH1, PMA1, and YEF3 was examined by ChIP in mog1Δ and wild‐type cells. We found that the gene presence of Rad6, Bre1, and Rtf1 was differentially affected by the loss of MOG1. The overall gene association of the three factors in mog1Δ cells was significantly reduced, but the effect was stronger at the 5'ORF (Figs 4A–C and EV5).
Figure 4. Mog1 affects the recruitment of Rad6, Bre1, and Rtf1 to 5'ORFs (5’) and 3'ORFs (3’) of genes.

-
AChIP analysis of TAP‐tagged Rad6 presence at ADH1, PMA1, and YEF3 coding regions (5’ and 3’) in WT or mog1Δ strains. Bar charts indicate the mean and standard deviation for at least three independent experiments. Significance of the differences was obtained using Student's t‐test (*P = 0.01–0.05; **P = 0.001–0.01; ***P = 0.0001–0.001; ****P < 0.0001).
-
B–DBre1‐TAP, Rtf1‐TAP, and Rpb3, respectively, were analyzed as described in (A).
-
EBar charts showing the relative chromatin occupancy to 5'ORFs (left panel) or 3'ORFs (right panel) of each factor relative to Rpb3 occupancy at these gene positions. Inputs were analyzed by Western blotting using anti‐TAP and anti‐PGK (loading control) antibodies to monitor protein levels (Fig EV5). Significance of the differences was obtained using Student's t‐test (*P = 0.01–0.05; **P = 0.001–0.01).
Figure EV5. Information about primers position and protein inputs levels used for ChIP analysis from Figures 2, 4 and 5.
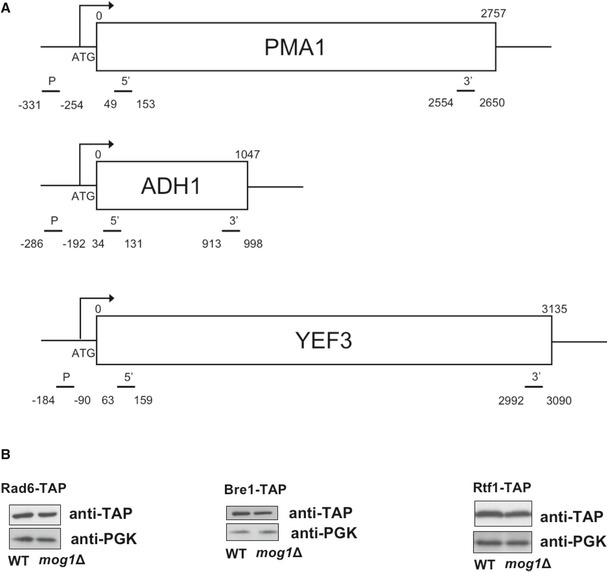
- Schematic representation of primer pairs used throughout this paper for ChIP analysis of PMA1, ADH1, and YEF3 genes.
- Inputs for ChIPs were analyzed by Western blotting using anti‐TAP and anti‐PGK (loading control) antibodies to monitor protein levels.
Rad6, Bre1, and Rtf1 are known to associate with RNAPII during transcription 8, 15. Therefore, their reduced gene recruitment could be due to a reduced RNAPII gene occupancy since the transcription rate is affected in mog1Δ cells (see Fig 3B and C). To clarify this matter, similar ChIP analyses of Rpb3 recruitment to the same three genes were performed in parallel to quantify Rpb3 association. As shown in Fig 4D, the relative occupancy of Rpb3 was affected by the loss of MOG1 at ADH1 and YEF3 with a stronger effect at 3'ORFs. Next, binding of Rpd3 was used to normalize Rad6, Bre1, and Rtf1 ChIP signals. Fig 4E shows that gene occupancy of these factors was differentially dependent on Rpb3 association. YEF3 gene occupancy of Rad6 at 3'ORFs or of Rtf1 at 5'ORFs was dependent on the presence of Rpb3, while Bre1 association depended on Rpb3 at both ORF regions. In contrast, the consequence of MOG1 deletion was independent of Rpb3 association at almost all PMA1 regions for all three factors.
Mog1 co‐precipitates with factors involved in transcription
Since the strongest observed effect of MOG1 deletion was a reduction in Rtf1 chromatin occupancy, we decided to assess a possible physical interaction between these two factors. Mog1 was therefore TAP‐tagged in a yeast strain expressing Rtf1‐PK. As shown in Fig 5A, immunoprecipitation of Mog1‐TAP co‐precipitated Rtf1‐PK suggesting that Mog1 could contact Rtf1 directly or indirectly. Such an interaction would be compatible with Mog1 association, at the least its transient association, with chromatin and its associated factors. To test this possibility, Mog1‐HA recruitment to PMA1 and YEF3 genes was studied using ChIP analysis. These experiments showed that Mog1‐HA gene occupancy was enhanced over control specifically at 5′ORFs (Fig 5B). The low, albeit statistically significant, absolute fold enhancement value is compatible with the possibility of a transient interaction between Mog1 and coding regions.
Figure 5. Mog1 co‐purifies with factors involved in transcription and associates with chromatin.
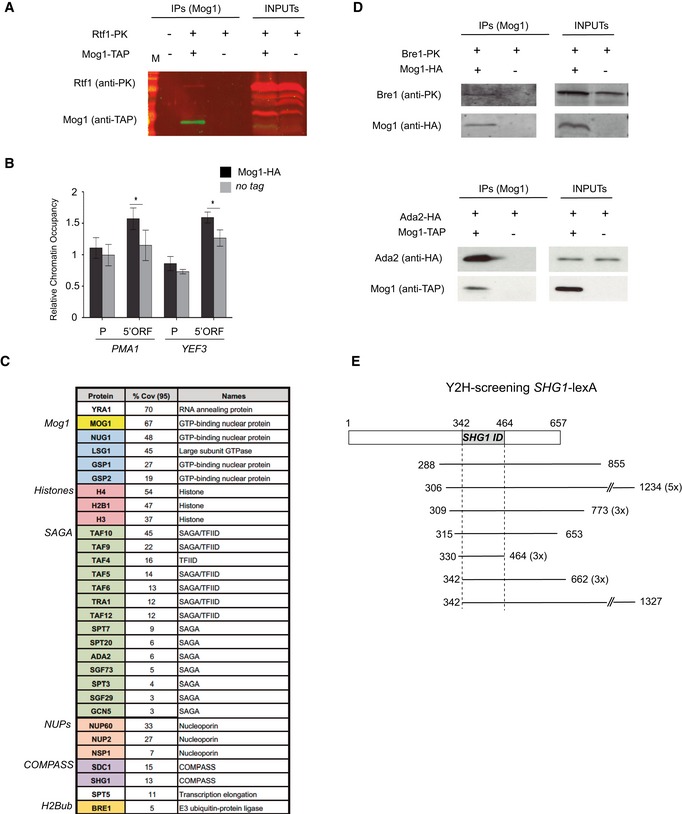
- Co‐purification of Rtf1‐PK in cells expressing Mog1‐TAP (+,+). A non‐tagged strain (−,−) and cells expressing only Rtf1‐PK (+,−) were included as negative controls for co‐purification. Inputs and IPs are depicted for each strain.
- ChIP analysis of the presence of HA‐tagged Mog1 compared to a non‐tagged strain (WT) at PMA1 and YEF3 promoter (P) and coding regions (5'ORF). The occupancy level was calculated as the signal ratio of the IP samples in relation to the input signal normalized to an intergenic region. Bar charts indicate the mean and standard deviation for at least three independent experiments. Significance of the differences was obtained using Student's t‐test and is presented as a P‐value (*P < 0.05).
- List of proteins identified from at least two independent LC‐MS/MS analyses of Mog1‐TAP purifications. Numbers indicate peptides of matching amino acids from identified peptides with a confidence greater or equal to 95% divided by the total number of amino acids in the sample (coverage 95%). An identifier of function or complex for each protein is listed.
- Mog1‐HA immunoprecipitation in cells expressing Bre1‐PK (upper panel) and Mog1‐TAP co‐precipitation in cells expressing Ada2‐HA (lower panel) are indicated as in (A) inputs and IPs are depicted for each strain.
- Representation of the different fragments of the MOG1 gene identified in genome‐wide Shg1 yeast two‐hybrid screening. The region common to all identified MOG1 clones between residues 114 and 154 is depicted in gray as the SHG1 ID (interaction domain). The start and end positions in the MOG1 gene are indicated as well as the number of clones containing this fragment.
Next, we decided to further identify all Mog1 co‐purifying proteins in an unbiased way. We therefore performed mass spectrometric analysis (LC/MS‐MS) of proteins that co‐purified with a TAP‐tagged version of Mog1 (compared to a non‐tagged strain) and focused our subsequent analysis on the co‐purification of other factors related to H2B ubiquitination, H3K4me3, and transcription. The mass spectrometric results confirmed that Mog1‐TAP specifically co‐purified with its previously described partners, the GTP‐binding proteins Gsp1, Gsp2, Nop1, and Lsg1 38, 40, which validated our Mog1‐TAP purification (Fig 5C and Dataset EV2). In addition, we identified the ubiquitin ligase Bre1, Spt5, fourteen subunits of the SAGA complex, and two subunits of the COMPASS (Set1C) complex, Shg1 and Sdc1. Several nucleoporins including Nup60, Nup2, and Nsp1 also co‐purified with Mog1‐TAP. To validate some of these interactions, we tagged Bre1 and Ada2 in cells expressing Mog1‐HA or Mog1‐TAP (see Materials and Methods). As shown in Fig 5D, Mog1 co‐precipitated with tagged Bre1 and Ada2. The interaction between COMPASS and Mog1 was validated through results obtained from a genome‐wide yeast two‐hybrid screen that was performed using SHG1 as the bait (Materials and Methods). Among the identified clones, 17 were found to contain fragments of the MOG1 gene (Table EV3; the complete results of this screen will be published elsewhere). The region common to all MOG1 clones was located between nts 342 and 464 (Fig 5E). The corresponding amino acid sequence between aa residues 114 and 154 may therefore demarcate a Shg1 interaction domain in Mog1. The combined data indicated that Mog1 interacts with several proteins that are involved in gene transcription.
Mog1 contributes to efficient mRNA nuclear export in coordination with the H2B ubiquitinating machinery
In addition to the SAGA DUBm genes, analyses of enriched gene ontologies (GO) showed that genes related to mRNA export were also overrepresented among the factors genetically interacting with mog1Δ (see Table EV1). Notably, several NUPs, including Nup60, Nup2, and Nsp1, which participate in mRNA transport, were identified as co‐purifying with Mog1‐TAP (Fig 5C), suggesting that Mog1 might be functionally involved in mRNA export. Furthermore, H2B monoubiquitination affects mRNP biogenesis and export 1, 2, 3. Controversial results have been published regarding a role for Mog1 in this process. While efficient nuclear export of mRNA occurred in mog1Δ cells of S. cerevisiae 38, such mRNA transport was defective in mog1Δ cells of S. pombe 59, 60. We decided to study the effect of mog1Δ on the cellular localization of poly (A)+ RNA under conditions that intensify mRNA export defects 18. These conditions consisted of increasing the temperature from 30°C to 39°C for 3 h, which results in a mRNA export block in some mutants that impede H2B monoubiquitination 61, 62. Figure 6 shows that mog1Δ led to inefficient mRNA export at 39°C when compared to the wild‐type control; however, it should be noted that the percentage of cells with such inefficient mRNA export was close to 30% and that the strength of this phenotype was clearly lower than that observed for the sus1Δ mutant, a bona fide export factor 34. We decided to further determine mRNA export in cells that combine MOG1 deletion with other mutations that affect H2B monoubiquitination. Deletion of BRE1 or LGE1 in a mog1Δ strain produces an additive effect on the blockage of mRNA export both in the intensity of the phenotype and in the percentage of cells affected (>90%; Fig 6). Notably, we observed that the phenotype of an rtf1Δ mutant showed mRNA accumulation in the nucleus as similar to a dot‐like pattern at 39°C and that this phenotype was greatly augmented by deletion of MOG1 (Fig 6 lowest panel). This result reinforced the strong functional interaction between these two factors. In summary, these data are consistent with a role for Mog1 in coordinating the monoubiquitination of H2B with mRNA export in yeast.
Figure 6. Mog1 contributes to efficient mRNA nuclear export in coordination with the H2B ubiquitination machinery.

Representative images of the analysis of nuclear mRNA export are shown. The cellular localization of poly(A)+ RNA was assessed by in situ hybridization using Cy3‐labeled oligo(dT) probes (red) in the indicated strains. In situ hybridization was monitored at 30°C and after shifting of the cells to 39°C for 3 h. DNA was stained by DAPI (blue). In all images, the scale bar corresponds to 2 μm. Images are representative pictures from two independent experiments of approximately 600 cells. The percentages indicate the number of cells showing mRNA export defect from these experiments.
Mog1 role in epigenetic regulation is independent of its ability to bind to Ran‐GTP
Overall, our results are supportive of a novel role for Mog1 in H2B monoubiquitination and H3K4 methylation, with the assumption that this function is independent of Mog1 participation in the nuclear protein import of these factors. Interestingly, previous work demonstrated that mutants such as MOG1‐E65K that impede Mog1 binding to Ran display a dramatic decrease in nuclear protein import 40. MOG1‐E65K cells grew more slowly than wild‐type cells at 37°C; however, their growth was not as slow as that of cells lacking MOG1 (see Fig 3B in 40). This observation suggested that Mog1 might perform additional functions in the cell that do not involve its interaction with Gsp1/Ran. For this purpose, we created the MOG1‐E65K point mutant and tested whether it could complement the double mutant mog1Δset1Δ in a similar manner to wild‐type MOG1. We found that mog1Δset1Δ cells expressing MOG1‐E65K grew in a similar manner to those expressing MOG1 (Fig 7A). This result suggested that binding to Ran was dispensable for this alternative Mog1 function. We also measured complementation of the levels of H3K4me3 and H2Bub1 in mog1Δ cells by MOG1‐E65K or MOG1 wild type, respectively. We found that the MOG1 deletion phenotype was largely compensated by Mog1 and by the Ran‐binding mutant MOG1‐E65K (Fig 7B). Furthermore, a specific role for Mog1 in H2B monoubiquitination was supported by the normal levels of this modification observed in cells lacking GSP2, a factor that interacts with Mog1 and is involved in nuclear protein import (Fig 7C). In summary, these results support a novel Ran independent role for Mog1 in modulating epigenetic modifications.
Figure 7. Mog1 binding to Ran is dispensable for its function in epigenetic modification.

- The Mog1E65K Ran‐binding mutant complements the slow growth of mog1ΔsetΔ mutants. Tenfold serial dilutions of wild‐type (WT), mog1Δ, set1Δ, and mog1ΔsetΔ strains transformed with wild‐type MOG1 (pMOG1) or with the Ran‐binding mutant (pMOG1E65K) as indicated were spotted on SC‐LEU and incubated for 2 days at the indicated temperatures.
- The Mog1E65K Ran‐binding mutant complements the low H3K4me3 and H2Bub1 levels of mog1Δ. The expression levels of H3K4me3, total H3, H2Bub1, total H2B, and PGK1 in whole‐cell lysates from wild‐type (WT) or from mog1Δ cells transformed with an empty plasmid (+pEmpty), wild‐type MOG1 (pMOG1), or a Ran‐binding mutant (pMOG1E65K) were analyzed by Western blotting using the corresponding antibodies.
- Gsp2 is not involved in H2Bub1 regulation. The expression levels of total H2B and of H2Bub1 were analyzed in whole‐cell lysates from wild‐type (WT) and gsp2Δ strains by Western blotting using anti‐total H2B and anti‐H2Bub1 antibodies.
Discussion
In this study, we discovered the following novel biological roles of Mog1.
Mog1 is required for H2Bub1‐ and RNAPII‐dependent transcription
The evolutionarily conserved protein Mog1 (human Ran guanine nucleotide release factor RANGNRF) was described as a coordinator of the Ran‐GTP cycle since it binds to Ran‐GTP and participates in the nuclear protein import pathway 38, 40. In humans, Mog1 regulates the function of the Nav1.5 cardiac sodium channel that is associated with the cardiac disease, Brugada syndrome (BrS) 63. Here, we identified a novel role for Mog1 in maintaining histone H2B monoubiquitination. The genetic interaction between MOG1 and the DUBm subunits Ubp8 and Sus1 26, 36, 44, drew our attention to a possible role for Mog1 in the same biological process. Surprisingly, the absence of MOG1 resulted in low levels of H2Bub1 independently of the presence of UBP8. Our experiments suggest that these low levels of H2Bub1 are a consequence of a reduced chromatin association of Rad6/Bre1 and Rtf1 mainly with the 5′ORF regions of highly transcribed genes. Rad6 association with chromatin has been proposed to occur in two steps that are modulated by interconnected mechanisms 8, 33. First, Rad6 accumulates at the promoter. Second, this accumulated Rad6 is then “handed off”, likely by Rtf1, to phosphorylated RNAPII through its association with the PAFc 64. Our ChIP experiments might indicate a role for Mog1 in this second step. We observed that the presence of all factors at the 5'ORFs and of Bre1 and Rtf1 at the 3'ORFs of all tested genes was reduced in the mog1Δ mutant. With the exception of the PMA1 gene, this reduction was also accompanied by a concomitant decrease in RNAPII (Rpb3) occupancy. Thus, it is possible that the overall reduction in H2Bub1 writers is a consequence of reduced RNAPII occupancy. However, the fact that Mog1 affects the association of Rad6 with gene bodies at 5'ORFs and of Rtf1 at 3'ORFs differentially, together with the finding that Mog1 and Rtf1 co‐precipitate might imply a more direct link between Mog1 and Rtf1. Recent research has proposed that Rtf1 can promote H2Bub1 levels through direct interaction with Rad6 15. It would be very interesting to determine whether Mog1 is involved in this interaction to better understand its mechanism of action in the maintenance of H2Bub1 levels.
Consistent with a reduction in the total levels of H2Bub1, deletion of MOG1 downregulates a 25% the global transcription rate (Fig 3D). This phenotype is not counterbalanced via stabilization of mRNA transcripts, since mRNA global abundance is decreased in almost 30% in a mog1Δ mutant, an effect not restricted to specific transcripts. At this stage, we cannot define whether the general drop in transcription in the absence of MOG1 reflects a direct or indirect effect of Mog1 in this process. In fact, deletion of MOG1 causes slow growth, a phenotype that correlates with a common gene expression signature highly similar to the environmental stress response (ESR) 65. However, gene set enrichment analyses (GSEA) of mog1Δ mutant did not find these gene ontologies, suggesting that mog1Δ gene expression profile is not a consequence of its slow growth rate (Table EV4). Furthermore, Mog1 physical interactions with components that are associated, in a broad sense, with transcription, together with Mog1 occupancy at gene bodies, might hint at a direct role. The fact that Mog1 directly affects the recruitment of Rtf1 to regions of specific genes could also explain this transcriptional drop. Future work will help to clarify this mechanism. Interestingly, Mog1 “knock‐down” experiments in zebrafish embryos or mutant MOG1 associated with Brugada syndrome patients significantly decreased the expression of several factors involved in cardiac morphogenesis 66. Our study might help to understand how Mog1 contributes to the downregulation of these genes in BrS patients bearing specific Mog1 mutations, which may possibly result from Mog1 affecting their H2Bub1 status.
Mog1 is required for the maintenance of H3K4me3 levels and interacts with the COMPASS subunit Shg1
The absence of MOG1 also led to decreased levels of H3K4me3. There are two possible explanations of this result: (i) Mog1 is required to sustain normal levels of H2Bub1, and, as a consequence, affects H3K4me3 levels indirectly, or, (ii) Mog1 modulates the crosstalk between H2Bub1 and H3K4me3 67. To discriminate between these two possibilities, we analyzed H3K4me2 levels and found that they were not affected by MOG1 deletion. This result suggested that Mog1 is not directly involved in establishing the crosstalk between H2Bub and H3K4me3, but that it rather affects H3K4me3 indirectly. Moreover, in contrast to H3K4me3, no differences in H3K79me3 or H3K36me3 levels were observed in the mog1Δ mutant in comparison with the wild‐type strain. These results were surprising but are comparable with previous observations of point mutants of the H2B C‐terminal such as H2BT122R and H2BR119D that reduced H2Bub1 and H3K4me3 levels without affecting H3K79me3 levels 49. These residues were mutated in the H2B C‐terminal have been shown to interact with Spp1 and to modulate Set1 function, thereby participating in H2B/H3K4 crosstalk. Notably, we also detected the interaction of two COMPASS subunits: Sdc1 and Shg1, with Mog1 in mass spectrometric analysis of Mog1‐TAP co‐purified proteins. Mog1‐Shg1 interaction was also detected in two‐hybrid experiments, which suggested that these two proteins could directly interact. Future analysis leading to the knowledge of how and when Mog1 might interact with Shg1, Bre1, Rtf1, and Ran is crucial in order to fully understand the role of Mog1 in regulating H2Bub1 levels and in transcription. Although our previous results using the Mog1‐E65K Ran‐binding mutant 40 suggest that the role of Mog1 in regulating H2Bub1 levels is Ran‐binding‐independent, we cannot exclude the possibility of an effect of Mog1 on regulator of chromosome condensation 1 (RCC1/RanGEF) binding to the nucleosome. RCC1 (yeast Prp20, 68) associated with chromatin generates a gradient of Ran‐GTP concentration that is required for protein transport 69, 70. The function of Mog1 in protein import is closely related to the action of RCC1 associated with chromatin. Strikingly, RCC1 directly associates with the nucleosome core particle (NCP) acidic patch where addition and removal of ubiquitin occur (see below; 70). Thus, it is possible that RCC1 directly or indirectly influences the mechanisms behind Rad6‐mediated ubiquitination. However, to our knowledge, no relationship between RCC1 and the ubiquitination of H2B has been predicted. In contrast, RCC1 has been linked to H2B phosphorylation in humans suggesting that this post‐translation modification affects RCC1 mobility and alters NF‐κB‐p65 transport 71. Further work is required to understand whether RCC1 contributes to the phenotypes shown here.
Mog1 is required for different cellular processes that require H2Bub1
Monoubiquitination of H2B is required for several fundamental processes in eukaryotic cells 50, 51. Among such processes, we found that mog1Δ cells, similarly to bre1Δ mog1Δ, are sensitive to replication stress caused by HU 52. Even though more work is required to elucidate if Mog1 is important for DNA replication, this preliminary result could indicate that the low levels of H2Bub1 observed in the mog1Δ cells have an impact on DNA replication.
Prior to mRNA nuclear export, there is tight control of the formation of export‐competent mRNPs 72. Previous research has shown that the post‐translation modifications of H2Bub1/H3K4me3 are required to couple some of the events of export‐competent mRNP formation with mRNA nuclear export 2. In agreement with deficiencies in these post‐translation modifications, mRNA export is partially affected in the absence of MOG1 (Fig 6). The role of Mog1 in mRNA transport is controversial 38, 59, 60. Clearly, Mog1 is not an export factor, but we were able to detect an enhancement of the accumulation of poly(A)+ RNA in the nucleus in the mog1Δ mutant by deleting BRE1, LGE1, or RTF1. Bre1 is known to partially block mRNA export upon cell treatment at 39°C 2, and this block was significantly augmented by the loss of MOG1. This finding suggests that both of these factors contribute to mRNA export. Notably, we also reported a novel phenotype for rtf1Δ cells that leads to the accumulation of poly(A)+ RNA into nuclear dot‐like structures (Fig 6). These dots became more intense and abundant in cells lacking both RTF1 and MOG1. These results, together with the ability of Rtf1 and Mog1 to co‐precipitate and the dependency of Rtf1 chromatin recruitment on Mog1, suggest a role for both factors in a common process.
Materials and Methods
Yeast strains, DNA recombinant work, and microbiological techniques
The yeast strains used in this study are listed in Table EV5. The quantitative polymerase chain reaction (qPCR) primers used are listed in Table EV6. Gene deletion and epitope tagging (TAP, GFP, and HA) were performed by homologous recombination. For growth analysis, yeast cultures were diluted to an OD600 of 0.5, and four serial dilutions were prepared (1:10; 1:100; 1:1,000; 1:10,000). Each dilution (5 μl) was spotted onto YP + Glucose or YP + Galactose plates and was incubated at indicated temperatures; generally, these were 30°C, 33°C, or 37°C.
Yeast cell extracts and western blot analysis
Yeast whole‐cell extracts (WCEs) were grown to an OD600 of 0.5–0.8 in YPD, and proteins were extracted following the trichloroacetic acid (TCA) method. WCEs, TAP‐containing complexes, and IPs were separated by gel electrophoresis (SDS–PAGE) and were then transferred onto nitrocellulose membranes (Amersham). Samples were analyzed by immunoblotting using specific antibodies (Table EV7) and detection reagents from ECL Amersham. In some cases, blots were developed using fluorescence detection on the Odyssey Infrared Imaging System from Licor Biosciences. In these cases, the secondary antibodies were from Licor Biosciences, catalog #926‐32211 (anti‐rabbit IgG IRDye800CW) or catalog #926‐68020 (anti‐mouse IgG IRDye 680LT). For signal quantification, at least three independent experiments were performed for each case. Error bars represent SD, and statistical difference is considered as *P = 0.01–0.05; **P = 0.001–0.01; and ***P < 0.001.
GFP localization
GFP‐tagged proteins were directly visualized in cells that were grown in SC medium to an OD600 of 0.3–0.5. The cells were washed with 1× PBS (13.7 mM NaCl, 0.27 mM KCl, 0.43 mM Na2HPO4, and 0.14 mM KH2PO4), centrifuged for 3 min at 2,000 rpm and were resuspended again in 1× PBS. Living cells tagged with GFP were visualized using a Leica TCS SP2 AOBS confocal microscope.
TAP‐tagged protein purification
The purification of TAP‐fusion proteins from yeast strains was performed by tandem affinity purification (TAP) as previously described 34. Briefly, TAP‐fusion proteins and their associated proteins were recovered from cell extracts by affinity selection on IgG beads. Next, these IgG beads were washed, and bound material was released by incubation with the tobacco etch virus (TEV) protease. This protease cleaves the TEV protease site in the TAP tag, thereby unveiling the calmodulin‐binding epitope in the fusion protein TAP tag and generating an eluate of the TAP‐tagged protein of interest termed the TEV eluate. Finally, the TAP‐tagged protein in the TEV eluate was purified by binding of the calmodulin‐binding epitope in the TAP tag to a Calmodulin Affinity Resin (Stratagene) in the presence of calcium. This purification step is required to remove both the TEV protease and traces of contaminants. The bound material was released with EGTA. This enriched protein fraction was termed the calmodulin eluate. After TCA precipitation, samples were subjected to Western blot analysis or to protein identification by mass spectrometry. LC‐MS/MS analyses were performed by the mass spectrometry service SCSIE at the University of Valencia following the standards of ProteoRed Spain (http://www.proteored.org) and http://www.scsie.uv.es.
Chromatin immunoprecipitation (ChIP)
Chromatin immunoprecipitation was performed as previously described [37,73] using 50 ml of yeast cultures grown to an OD600 of 0.5 in YP + Glucose. Cultures were cross‐linked for 20 min at room temperature with formaldehyde (1% final concentration; Sigma) and were later quenched with 125 mM glycine. Subsequently, the cells were collected by centrifugation and washed three times with 25 ml of cold Tris‐saline buffer (150 mM NaCl, 20 mM Tris–HCl, pH 7.5). The cells were broken by adding 300 μl of lysis buffer (50 mM HEPES‐KOH at pH 7.5, 140 mM NaCl, 1 mM EDTA, 1 mM PMSF, and proteases inhibitors) and 200 μl glass beads and incubating for 15 min at 4°C. Cell extracts were sonicated for 30 min in a Bioruptor sonicator (Diagenode) using 30‐s on/30‐s off cycles. An aliquot (10 μl) of the chromatin extract was saved as the input. The rest of the lysate was used for immunoprecipitation by incubating with the required specific antibodies (Table EV7) and magnetic beads (Dynabeads, Invitrogen) for 2 h at 4°C. The beads were subsequently washed with the following buffers: twice with lysis buffer, twice with lysis buffer supplemented with 360 mM NaCl, twice with wash buffer (10 mM Tris–HCl, pH 8.0, 250 mM LiCl, 0.5% NP‐40, 125 mM Nadeoxycholol, and 1 mM EDTA), and once with TE buffer. Samples were eluted by adding 50 μl of elution buffer (50 mM Tris–HCl, pH 8, 10 mM EDTA, 1% SDS) to the beads and incubating for 10 min at 65°C. This step was repeated twice. Input and immunoprecipitation (IP) samples were incubated overnight at 65°C to reverse the cross‐linking, following which 100 μg/250 μl proteinase K (Ambion) was incubated with the IP and input for 1.5 h at 45°C. DNA was extracted using a DNA Purification Kit (Zymo Research) following the manufacturer's instructions. This DNA was analyzed using qPCR with specific primers (Table EV6). In all cases, at least three biological replicates were performed to calculate the standard deviation.
Immunoprecipitations and protein purifications
For protein purifications and immunoprecipitation (IP), 50 ml of yeast cultures grown to an OD600 of 0.5 in YP + Glucose was collected. The cells were washed once with 1× LB buffer (10 mM Tris–HCl pH 8.0, 150 mM NaCl, 20% glycerol (v/v), and Nonidet P‐40 0.1% (v/v)) and were then resuspended in lysis buffer (LB 1×, 0.5 mM DTT, 1 mM PMSF and 1× complete protease cocktail inhibitor (Roche)). An equal volume of glass beads was added, and the cells were broken by four pulses of vortexing over 1 min at 4°C. Following centrifugation, the cell supernatant was incubated directly with magnetic beads (Dynabeads, Invitrogen) for TAP‐tagged proteins or with magnetic beads precoated with the specific antibody (anti‐PK) for 2 h at 4°C in a turning wheel. The immunoprecipitated samples were washed with three 10‐min washes with 1× LB and 0.5 mM DTT and were resuspended in 50 μl of sodium dodecyl sulfate–polyacrylamide (SDS–PAGE) sample buffer. Samples were then analyzed by SDS–PAGE and Western blotting.
Yeast two‐hybrid screening
Yeast two‐hybrid screening was carried out by Hybrigenics, SA, Paris. A mating strategy was employed, using a genomic library introduced in Y187 cells and L40∆G cells producing a C‐terminal fusion of LexA to the SHG1 (YBR258C) open reading frame 74. A total of 123 positive clones were identified and analyzed as described 74; 17 of these clones contained fragments of the MOG1 gene. The entire results of the screen will be published elsewhere (B. Dichtl, unpublished results).
Fluorescence in situ hybridization (FISH)
Fluorescence in situ hybridization for detection of poly(A)+ RNA was performed by growing yeast cells in 100 ml of YP + Glucose medium overnight at 30°C to an OD600 of 0.5 and then shifting the cells to 39°C for 3 h. Subsequently, the cells were fixed by adding 4% formaldehyde and were incubated for 60 min at room temperature. Formaldehyde was removed by two rounds of centrifugation and washing with 0.1 M potassium phosphate (pH 6.4). The cells were resuspended in ice‐cold washing buffer (1.2 M sorbitol and 0.1 M potassium phosphate, pH 6.4), digested with 0.5 mg/ml of Zymolyase 100T, and applied onto poly‐L‐lysine‐coated slide wells. Cells not adhered to the slides were removed by aspiration. Cells were rehydrated with 2× SSC (0.15 M NaCl and 0.015 M sodium citrate) and were hybridized overnight at 37°C in 20 μl of prehybridization buffer containing 0.8 pmol of Cy3 end‐labeled oligo (dT) in a humid chamber. Following this hybridization, the slides were washed with 2× SSC at room temperature for 5 min and after with 1× SSC. Then, they were air‐dried and mounted using 3.5 μl/well of VECTASHIELD® Mounting Medium containing DAPI. Cy3‐oligo (dT) fluorescence was detected using a Leica TCS SP2 AOBS confocal microscope. Two biological replicates were performed for all experiments. The selected images in Fig 6 are representative of three different pictures containing more than 600 cells per experiment.
Genomic Run‐On (GRO) and measurement of mRNA levels
Genomic Run‐On experiments were performed in triplicate in custom made macroarrays as described in 54 with the changes described in 75. Normalization between samples was done by using three different B. subtilis genes that were in vitro‐transcribed from linearized plasmids (Lys, Thr, and Phe in a pGIBS plasmid: ATCC clones 87,482, 87,483, and 87,484). Equal amounts of a mixture of these three transcripts were added to the yeast cell samples before RNA extraction to be used as internal spike‐in controls similarly to the protocol described in 76. mRNA synthesis rates (SR) were calculated from raw GRO data that were bioinformatically processed as described in 75 and then divided by the relative median cell volume of each strain as determined with a Coulter counter device 77. The total RNAPII synthesis rate was determined by using the median of the distribution of the whole dataset of individual synthesis rates for all genes.
mRNA levels (RA) were determined using aliquots of the same samples used for GRO as described in 54 and 75. In this experiment, the total RNA was extracted and quantified for the same number of cells. The proportion of mRNA per total RNA was calculated by a dot‐blot procedure using 32P‐labeled oligo dT hybridization as previously described 75. The level of total mRNA per cell was calculated from the total RNA/cell and the proportion of mRNA/total RNA. The relative change in total mRNA/cell in the mog1Δ mutant vs. the WT was used to normalize the whole set of individual mRNA levels.
mRNA half‐lives were determined by dividing RA data by SR data as described previously 54.
Accession numbers
The Gene Expression Omnibus (GEO) accession number for genomic data is GSE109435.
Author contributions
SR‐N designed the study; PO‐C, JS‐Q, and CN‐C performed most of the experiments; PO‐C, MEP‐M, and JEP‐O performed and analyzed the GRO experiments; BD performed and analyzed the yeast 2‐hybrid experiments; SB and LMS contributed with suggestions and interpretations of the results; and PO‐C, JEP‐O, LMS, SB, and SR‐N wrote the manuscript.
Conflict of interest
The authors declare that they have no conflict of interest.
Supporting information
Expanded View Figures PDF
Dataset EV1
Dataset EV2
Table EV1
Table EV2
Table EV3
Table EV4
Table EV5
Table EV6
Table EV7
Review Process File
Acknowledgments
We are grateful to Dr. J. Fassler (The University of Iowa) for Mog1 plasmids and to Mª Eugenia Gas and Lucía Casares from the Rodríguez‐Navarro laboratory for their experimental support. Special thanks to Dr Martínez‐Beneyto for her contribution in creating the synopsis image. PO‐C, JS‐Q and CN‐C were supported by the FPI, FPU, and PROMETEO (BES‐2012058587, FPU15/03862, PROMETEO 2016/093). This study was supported by funds to SR‐N from the Spanish MINECO (BFU2011‐23418, BFU2014‐57636) and the Generalitat Valenciana (PROMETEO 2012/061 and ACOMP2014/061) and to JEP‐O (BFU2016‐77728‐C3‐3‐P, PROMETEOII 2015/006).
EMBO Reports (2018) 19: e45992
References
- 1. Rodríguez‐Navarro S, Hurt E (2011) Linking gene regulation to mRNA production and export. Curr Opin Cell Biol 23: 302–309 [DOI] [PubMed] [Google Scholar]
- 2. Vitaliano‐Prunier A, Babour A, Hérissant L, Apponi L, Margaritis T, Holstege FCP, Corbett AH, Gwizdek C, Dargemont C (2012) H2B ubiquitylation controls the formation of export‐competent mRNP. Mol Cell 45: 132–139 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Hérissant L, Moehle EA, Bertaccini D, Van Dorsselaer A, Schaeffer‐Reiss C, Guthrie C, Dargemont C (2014) H2B ubiquitylation modulates spliceosome assembly and function in budding yeast. Biol Cell 106: 126–138 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Osley MA (2006) Regulation of histone H2A and H2B ubiquitylation. Brief Funct Genomic Proteomic 5: 179–189 [DOI] [PubMed] [Google Scholar]
- 5. Pavri R, Zhu B, Li G, Trojer P, Mandal S, Shilatifard A, Reinberg D (2006) Histone H2B monoubiquitination functions cooperatively with FACT to regulate elongation by RNA polymerase II. Cell 125: 703–717 [DOI] [PubMed] [Google Scholar]
- 6. Bhaumik SR, Smith E, Shilatifard A (2007) Covalent modifications of histones during development and disease pathogenesis. Nat Struct Mol Biol 14: 1008–1016 [DOI] [PubMed] [Google Scholar]
- 7. Cole AJ, Clifton‐Bligh R, Marsh DJ (2015) Histone H2B monoubiquitination: roles to play in human malignancy. Endocr Relat Cancer 22: T19–T33 [DOI] [PubMed] [Google Scholar]
- 8. Wood A, Krogan NJ, Dover J, Schneider J, Heidt J, Boateng MA, Dean K, Golshani A, Zhang Y, Greenblatt JF et al (2003) Bre1, an E3 ubiquitin ligase required for recruitment and substrate selection of Rad6 at a promoter. Mol Cell 11: 267–274 [DOI] [PubMed] [Google Scholar]
- 9. Song Y‐H, Ahn SH (2010) A Bre1‐associated protein, large 1 (Lge1), promotes H2B ubiquitylation during the early stages of transcription elongation. J Biol Chem 285: 2361–2367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Kim J, Guermah M, McGinty RK, Lee J‐S, Tang Z, Milne TA, Shilatifard A, Muir TW, Roeder RG (2009) RAD6‐mediated transcription‐coupled H2B ubiquitylation directly stimulates H3K4 methylation in human cells. Cell 137: 459–471 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Hwang WW, Venkatasubrahmanyam S, Ianculescu AG, Tong A, Boone C, Madhani HD (2003) A conserved RING finger protein required for histone H2B monoubiquitination and cell size control. Mol Cell 11: 261–266 [DOI] [PubMed] [Google Scholar]
- 12. Weake VM, Workman JL (2008) Clearing the way for unpaused polymerases. Cell 134: 16–18 [DOI] [PubMed] [Google Scholar]
- 13. Van Oss SB, Cucinotta CE, Arndt KM (2017) Emerging insights into the roles of the Paf1 complex in gene regulation. Trends Biochem Sci 42: 788–798 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Fuchs G, Oren M (2014) Writing and reading H2B monoubiquitylation. Biochim Biophys Acta 1839: 694–701 [DOI] [PubMed] [Google Scholar]
- 15. Van Oss SB, Shirra MK, Bataille AR, Wier AD, Yen K, Vinayachandran V, Byeon I‐JL, Cucinotta CE, Héroux A, Jeon J et al (2016) The histone modification domain of Paf1 complex subunit Rtf1 directly stimulates H2B ubiquitylation through an interaction with Rad6. Mol Cell 64: 1–12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Dehé P‐M, Géli V (2006) The multiple faces of Set1. Biochem Cell Biol 84: 536–548 [DOI] [PubMed] [Google Scholar]
- 17. Shahbazian M, Zhang K, Grunstein M (2005) Histone H2B ubiquitylation controls processive methylation but not monomethylation by Dot1 and Set1. Mol Cell 19: 271–277 [DOI] [PubMed] [Google Scholar]
- 18. Vitaliano‐Prunier A, Menant A, Hobeika M, Géli V, Gwizdek C, Dargemont C (2008) Ubiquitylation of the COMPASS component Swd2 links H2B ubiquitylation to H3K4 trimethylation. Nat Cell Biol 10: 1365–1371 [DOI] [PubMed] [Google Scholar]
- 19. Soares LM, Buratowski S (2012) Yeast Swd2 is essential because of antagonism between Set1 histone methyltransferase complex and APT (associated with Pta1) termination factor. J Biol Chem 287: 15219–15231 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Shilatifard A (2006) Chromatin modifications by methylation and ubiquitination: implications in the regulation of gene expression. Annu Rev Biochem 75: 243–269 [DOI] [PubMed] [Google Scholar]
- 21. Atanassov BS, Mohan RD, Lan X, Kuang X, Lu Y, Lin K, McIvor E, Li W, Zhang Y, Florens L et al (2016) ATXN7L3 and ENY2 coordinate activity of multiple H2B deubiquitinases important for cellular proliferation and tumor growth. Mol Cell 62: 558–571 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Daniel JA, Torok MS, Sun Z‐W, Schieltz D, Allis CD, Yates JR, Grant PA (2004) Deubiquitination of histone H2B by a yeast acetyltransferase complex regulates transcription. J Biol Chem 279: 1867–1871 [DOI] [PubMed] [Google Scholar]
- 23. Galan A, Rodríguez‐Navarro S (2012) Sus1/ENY2: a multitasking protein in eukaryotic gene expression. Crit Rev Biochem Mol Biol 47: 556–568 [DOI] [PubMed] [Google Scholar]
- 24. Helmlinger D, Hardy S, Abou‐Sleymane G, Eberlin A, Bowman AB, Gansmüller A, Picaud S, Zoghbi HY, Trottier Y, Tora L et al (2006) Glutamine‐expanded ataxin‐7 alters TFTC/STAGA recruitment and chromatin structure leading to photoreceptor dysfunction. PLoS Biol 4: e67 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Henry KW, Wyce A, Lo W‐S, Duggan LJ, Emre NCT, Kao C‐F, Pillus L, Shilatifard A, Osley MA, Berger SL (2003) Transcriptional activation via sequential histone H2B ubiquitylation and deubiquitylation, mediated by SAGA‐associated Ubp8. Genes Dev 17: 2648–2663 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Ingvarsdottir K, Krogan NJ, Emre NCT, Wyce A, Thompson NJ, Emili A, Hughes TR, Greenblatt JF, Berger SL (2005) H2B ubiquitin protease Ubp8 and Sgf11 constitute a discrete functional module within the Saccharomyces cerevisiae SAGA complex. Mol Cell Biol 25: 1162–1172 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Köhler A, Schneider M, Cabal GG, Nehrbass U, Hurt E (2008) Yeast Ataxin‐7 links histone deubiquitination with gene gating and mRNA export. Nat Cell Biol 10: 707–715 [DOI] [PubMed] [Google Scholar]
- 28. Köhler A, Zimmerman E, Schneider M, Hurt E, Zheng N (2010) Structural basis for assembly and activation of the heterotetrameric SAGA histone H2B deubiquitinase module. Cell 141: 606–617 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Lee K, Florens L, Swanson S, Washburn M, Workman J (2005) The deubiquitylation activity of Ubp8 is dependent upon Sgf11 and its association with the SAGA complex. Mol Cell Biol 25: 1173–1182 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Lee KK, Swanson SK, Florens L, Washburn MP, Workman JL (2009) Yeast Sgf73/Ataxin‐7 serves to anchor the deubiquitination module into both SAGA and Slik(SALSA) HAT complexes. Epigenetics Chromatin 2: 2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Samara NL, Datta AB, Berndsen CE, Zhang X, Yao T, Cohen RE, Wolberger C (2010) Structural insights into the assembly and function of the SAGA deubiquitinating module. Science 328: 1025–1029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Morgan MT, Haj‐Yahya M, Ringel AE, Bandi P, Brik A, Wolberger C (2016) Structural basis for histone H2B deubiquitination by the SAGA DUB module. Science 351: 725–728 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Gallego LD, Ghodgaonkar Steger M, Polyansky AA, Schubert T, Zagrovic B, Zheng N, Clausen T, Herzog F, Köhler A (2016) Structural mechanism for the recognition and ubiquitination of a single nucleosome residue by Rad6‐Bre1. Proc Natl Acad Sci USA 113: 10553–10558 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Rodríguez‐Navarro S, Fischer T, Luo M‐J, Antúnez O, Brettschneider S, Lechner J, Pérez‐Ortín JE, Reed R, Hurt E (2004) Sus1, a functional component of the SAGA histone acetylase complex and the nuclear pore‐associated mRNA export machinery. Cell 116: 75–86 [DOI] [PubMed] [Google Scholar]
- 35. Pascual‐García P, Govind CK, Queralt E, Cuenca‐Bono B, Llopis A, Chavez S, Hinnebusch AG, Rodríguez‐Navarro S (2008) Sus1 is recruited to coding regions and functions during transcription elongation in association with SAGA and TREX2. Genes Dev 22: 2811–2822 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Köhler A, Pascual‐García P, Llopis A, Zapater M, Posas F, Hurt E, Rodríguez‐Navarro S (2006) The mRNA export factor Sus1 is involved in Spt/Ada/Gcn5 acetyltransferase‐mediated H2B deubiquitinylation through its interaction with Ubp8 and Sgf11. Mol Biol Cell 17: 4228–4236 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. García‐Oliver E, Pascual‐García P, García‐Molinero V, Lenstra TL, Holstege FCP, Rodríguez‐Navarro S (2013) A novel role for Sem1 and TREX‐2 in transcription involves their impact on recruitment and H2B deubiquitylation activity of SAGA. Nucleic Acids Res 41: 5655–5668 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Oki M, Nishimoto T (1998) A protein required for nuclear‐protein import, Mog1p, directly interacts with GTP‐Gsp1p, the Saccharomyces cerevisiae ran homologue. Proc Natl Acad Sci USA 95: 15388–15393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Steggerda SM, Black BE, Paschal BM (2000) Monoclonal antibodies to NTF2 inhibit nuclear protein import by preventing nuclear translocation of the GTPase Ran. Mol Biol Cell 11: 703–719 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Baker RP, Harreman MT, Eccleston JF, Corbett AH, Stewart M (2001) Interaction between Ran and Mog1 is required for efficient nuclear protein import. J Biol Chem 276: 41255–41262 [DOI] [PubMed] [Google Scholar]
- 41. Baryshnikova A, Costanzo M, Kim Y, Ding H, Koh J, Toufighi K, Youn J‐Y, Ou J, San Luis B‐J, Bandyopadhyay S et al (2010) Quantitative analysis of fitness and genetic interactions in yeast on a genome scale. Nat Methods 7: 1017–1024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Costanzo M, Baryshnikova A, Bellay J, Kim Y, Spear ED, Sevier CS, Ding H, Koh JLY, Toufighi K, Mostafavi S et al (2010) The genetic landscape of a cell. Science 327: 425–431 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Wood A, Schneider J, Dover J, Johnston M, Shilatifard A (2003) The Paf1 complex is essential for histone monoubiquitination by the Rad6‐Bre1 complex, which signals for histone methylation by COMPASS and Dot1p. J Biol Chem 278: 34739–34742 [DOI] [PubMed] [Google Scholar]
- 44. Shukla A, Stanojevic N, Duan Z, Shadle T, Bhaumik SR (2006) Functional analysis of H2B‐Lys‐123 ubiquitination in regulation of H3‐Lys‐4 methylation and recruitment of RNA polymerase II at the coding sequences of several active genes in vivo . J Biol Chem 281: 19045–19054 [DOI] [PubMed] [Google Scholar]
- 45. Shilatifard A (2012) The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu Rev Biochem 81: 65–95 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Schneider J, Wood A, Lee J‐S, Schuster R, Dueker J, Maguire C, Swanson SK, Florens L, Washburn MP, Shilatifard A (2005) Molecular regulation of histone H3 trimethylation by COMPASS and the regulation of gene expression. Mol Cell 19: 849–856 [DOI] [PubMed] [Google Scholar]
- 47. Ng HH, Robert F, Young RA, Struhl K (2003) Targeted recruitment of Set1 histone methylase by elongating Pol II provides a localized mark and memory of recent transcriptional activity. Mol Cell 11: 709–719 [DOI] [PubMed] [Google Scholar]
- 48. Vlaming H, van Leeuwen F (2016) The upstreams and downstreams of H3K79 methylation by DOT1L. Chromosoma 125: 593–605 [DOI] [PubMed] [Google Scholar]
- 49. Chandrasekharan MB, Huang F, Chen Y‐C, Sun Z‐W (2010) Histone H2B C‐terminal helix mediates trans‐histone H3K4 methylation independent of H2B ubiquitination. Mol Cell Biol 30: 3216–3232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Zhang W, Yeung CHL, Wu L, Yuen KWY (2017) E3 ubiquitin ligase Bre1 couples sister chromatid cohesion establishment to DNA replication in Saccharomyces cerevisiae . eLife 6: 1082 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Bonnet J, Devys D, Tora L (2014) Histone H2B ubiquitination: signaling not scrapping. Drug Discov Today Technol 12: e19–e27 [DOI] [PubMed] [Google Scholar]
- 52. Trujillo KM, Osley MA (2012) A role for H2B ubiquitylation in DNA replication. Mol Cell 48: 734–746 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Zhang Y (2003) Transcriptional regulation by histone ubiquitination and deubiquitination. Genes Dev 17: 2733–2740 [DOI] [PubMed] [Google Scholar]
- 54. García‐Martínez J, Aranda A, Pérez‐Ortín JE (2004) Genomic run‐on evaluates transcription rates for all yeast genes and identifies gene regulatory mechanisms. Mol Cell 15: 303–313 [DOI] [PubMed] [Google Scholar]
- 55. Huisinga KL, Pugh BF (2004) A genome‐wide housekeeping role for TFIID and a highly regulated stress‐related role for SAGA in Saccharomyces cerevisiae . Mol Cell 13: 573–585 [DOI] [PubMed] [Google Scholar]
- 56. Baptista T, Grünberg S, Minoungou N, Koster MJE, Timmers HTM, Hahn S, Devys D, Tora L (2017) SAGA is a general cofactor for RNA polymerase II transcription. Mol Cell 68: 130–143.e135 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Warfield L, Ramachandran S, Baptista T, Devys D, Tora L, Hahn S (2017) Transcription of nearly all yeast RNA polymerase II‐transcribed genes is dependent on transcription factor TFIID. Mol Cell 68: 118–129.e5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. García‐Molinero V, García‐Martínez J, Reja R, Furió‐Tarí P, Antúnez O, Vinayachandran V, Conesa A, Pugh BF, Pérez‐Ortín JE, Rodríguez‐Navarro S (2018) The SAGA/TREX‐2 subunit Sus1 binds widely to transcribed genes and affects mRNA turnover globally. Epigenetics Chromatin 11: 13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Oki M, Ma L, Wang Y, Hatanaka A, Miyazato C, Tatebayashi K, Nishitani H, Uchida H, Nishimoto T (2007) Identification of novel suppressors for Mog1 implies its involvement in RNA metabolism, lipid metabolism and signal transduction. Gene 400: 114–121 [DOI] [PubMed] [Google Scholar]
- 60. Tatebayashi K, Tani T, Ikeda H (2001) Fission yeast Mog1p homologue, which interacts with the small GTPase Ran, is required for mitosis‐to‐interphase transition and poly(A)(+) RNA metabolism. Genetics 157: 1513–1522 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Gwizdek C, Iglesias N, Rodriguez MS, Ossareh‐Nazari B, Hobeika M, Divita G, Stutz F, Dargemont C (2006) Ubiquitin‐associated domain of Mex67 synchronizes recruitment of the mRNA export machinery with transcription. Proc Natl Acad Sci USA 103: 16376–16381 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Iglesias N, Tutucci E, Gwizdek C, Vinciguerra P, Von Dach E, Corbett AH, Dargemont C, Stutz F (2010) Ubiquitin‐mediated mRNP dynamics and surveillance prior to budding yeast mRNA export. Genes Dev 24: 1927–1938 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Kattygnarath D, Maugenre S, Neyroud N, Balse E, Ichai C, Denjoy I, Dilanian G, Martins RP, Fressart V, Berthet M et al (2011) MOG1: a new susceptibility gene for Brugada syndrome. Circ Cardiovasc Genet 4: 261–268 [DOI] [PubMed] [Google Scholar]
- 64. Xiao T, Kao C‐F, Krogan NJ, Sun Z‐W, Greenblatt JF, Osley MA, Strahl BD (2005) Histone H2B ubiquitylation is associated with elongating RNA polymerase II. Mol Cell Biol 25: 637–651 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. O'Duibhir E, Lijnzaad P, Benschop JJ, Lenstra TL, Van Leenen D, Groot Koerkamp MJA, Margaritis T, Brok MO, Kemmeren P, Holstege FCP (2014) Cell cycle population effects in perturbation studies. Mol Syst Biol 10: 732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Zhou J, Wang L, Zuo M, Wang X, Ahmed ASI, Chen Q, Wang QK (2016) Cardiac sodium channel regulator MOG1 regulates cardiac morphogenesis and rhythm. Sci Rep 6: 1–12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Sun Z‐W, Allis CD (2002) Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature 418: 104–108 [DOI] [PubMed] [Google Scholar]
- 68. Lee A, Tam R, Belhumeur P, DiPaolo T, Clark MW (1993) Prp20, the Saccharomyces cerevisiae homolog of the regulator of chromosome condensation, RCC1, interacts with double‐stranded DNA through a multi‐component complex containing GTP‐binding proteins. J Cell Sci 106(Pt 1): 287–298 [DOI] [PubMed] [Google Scholar]
- 69. Carazo‐Salas RE, Guarguaglini G, Gruss OJ, Segref A, Karsenti E, Mattaj IW (1999) Generation of GTP‐bound Ran by RCC1 is required for chromatin‐induced mitotic spindle formation. Nature 400: 178–181 [DOI] [PubMed] [Google Scholar]
- 70. Kalashnikova AA, Porter‐Goff ME, Muthurajan UM, Luger K, Hansen JC (2013) The role of the nucleosome acidic patch in modulating higher order chromatin structure. J R Soc Interface 10: 20121022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Wong C‐H, Chan H, Ho C‐Y, Lai S‐K, Chan K‐S, Koh C‐G, Li H‐Y (2008) Apoptotic histone modification inhibits nuclear transport by regulating RCC1. Nat Cell Biol 11: 36–45 [DOI] [PubMed] [Google Scholar]
- 72. Mitchell SF, Parker R (2014) Principles and properties of eukaryotic mRNPs. Mol Cell 54: 547–558 [DOI] [PubMed] [Google Scholar]
- 73. Pamblanco M, Oliete‐Calvo P, García‐Oliver E, Luz Valero M, Sanchez del Pino MM, Rodríguez‐Navarro S (2014) Unveiling novel interactions of histone chaperone Asf1 linked to TREX‐2 factors Sus1 and Thp1. Nucleus 5: 247–259 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Fromont‐Racine M, Rain JC, Legrain P (1997) Toward a functional analysis of the yeast genome through exhaustive two‐hybrid screens. Nat Genet 16: 277–282 [DOI] [PubMed] [Google Scholar]
- 75. Canadell D, García‐Martínez J, Alepuz P, Pérez‐Ortín JE, Ariño J (2015) Impact of high pH stress on yeast gene expression: a comprehensive analysis of mRNA turnover during stress responses. Biochim Biophys Acta 1849: 653–664 [DOI] [PubMed] [Google Scholar]
- 76. van de Peppel J, Kemmeren P, van Bakel H, Radonjic M, Van Leenen D, Holstege FCP (2003) Monitoring global messenger RNA changes in externally controlled microarray experiments. EMBO Rep 4: 387–393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. García‐Martínez J, Troulé K, Chavez S, Pérez‐Ortín JE (2016) Growth rate controls mRNA turnover in steady and non‐steady states. RNA Biol 13: 1175–1181 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Expanded View Figures PDF
Dataset EV1
Dataset EV2
Table EV1
Table EV2
Table EV3
Table EV4
Table EV5
Table EV6
Table EV7
Review Process File


