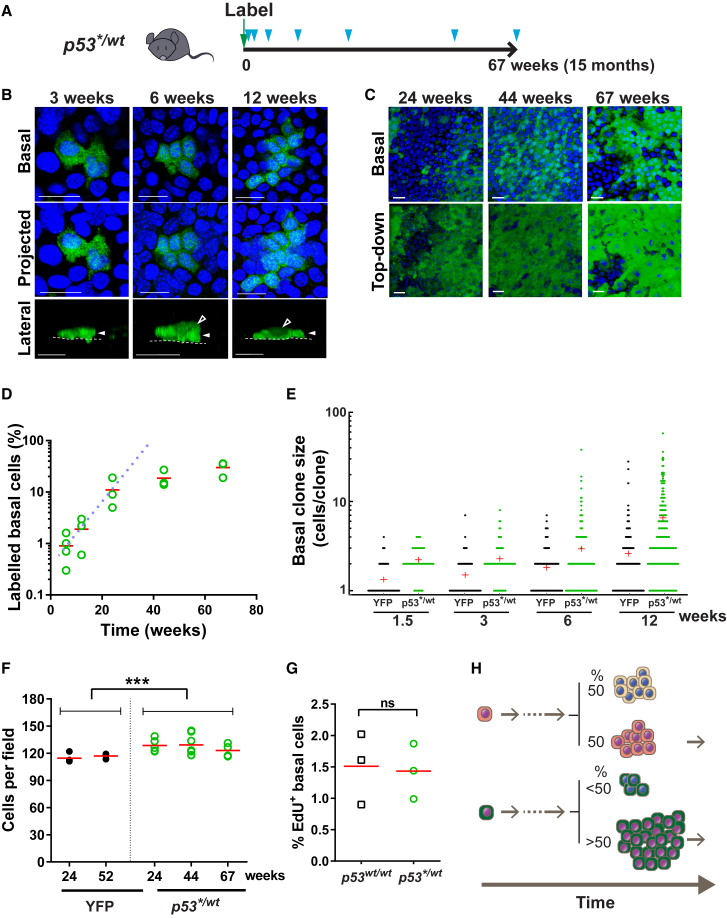
Figure 3.
Heterozygous p53R245W (p53∗/wt) Mutant Cell Fate
(A) Protocol: clonal labeling of AhcreERTp53flR245W/wt (p53∗/wt) mice followed by sampling (triangles).
(B) Rendered confocal z stacks showing typical p53∗/wt clones in back skin epidermis. Basal, top-down view of basal layer; projected, top-down view through all nucleated cell layers; lateral, side view. Cells in basal and first suprabasal cell layers are indicated by closed and open arrowheads, respectively; dotted line, basement membrane. Green, GFP; blue, DAPI. Scale bars, 20 μm.
(C) Rendered z stacks showing basal and top-down views of typical whole mounts at times indicated. Green, GFP; blue, DAPI. Scale bars, 20 μm.
(D) Proportion of labeled basal cells at indicated time points. Averaged value from 4–5 fields per animal, n = 3 animals per time point except n = 4 animals at the 6-week time point. Red lines, mean value. Dotted line indicates the expected growth of p53∗/wt basal cells clone area if the rate of expansion is constant.
(E) Clone size distributions (basal cells per clone). Red cross indicates mean clone size. Data are from 3–5 mice per time point. For p53∗/wt and RYFP, respectively, n = 50, 93 clones at 1.5 weeks, 75, 106 clones at 3 weeks, 199, 181 clones at 6 weeks and 196, 183 clones at 12 weeks.
(F) Mean basal cell density in p53∗/wt and RYFP control mice. Data are means of 5 fields per mouse. n ≥ 4 mice at each time point/group. Exact sample numbers are in corresponding supplementary table, ∗∗∗p = 0.0007 by Mann-Whitney test.
(G) Average percentage of EdU-labeled basal cells in p53∗/wt clones at the 12-week time point compared to non-labeled cells (p53wt/wt) in same mouse. n = 3 mice. ns, no significant difference by paired t test.
(H) Schematic illustration of p53∗/wt cell behavior. A bias in the fate of p53∗/wt cell can result in an increase in the proportion of p53∗/wt progenitors in the IFE, even if the rate of mutant cell division was the same as that of wild-type cells.
See also Figure S1–S6 and Tables S2 and S7.