Figure 2.
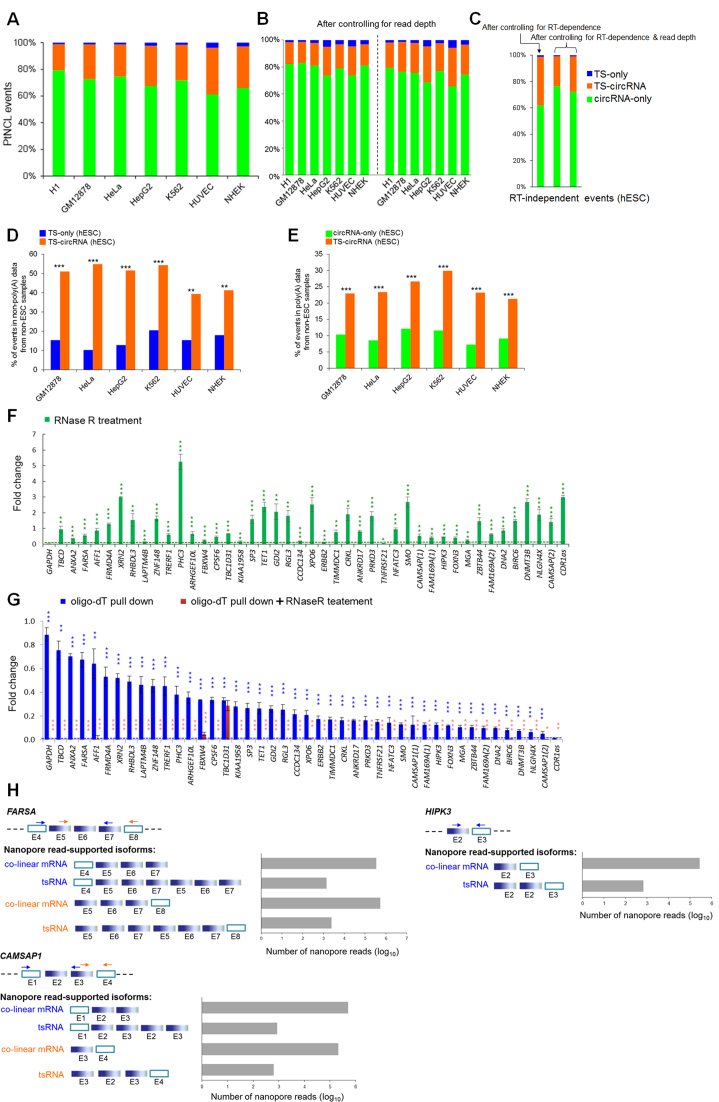
Different types of NCL events. (A–C) Distribution of three types of NCL events: TS-only, circRNA-only and TS-circRNA events before (A) and after (B and C) controlling for read depth/RT-dependence. For (B and C), read depth was normalized by randomly selecting an equal number (60 million, left; 120 million, right) of reads from both poly(A)- and non-poly(A)-selected data for the examined cell lines (Supplementary Tables S2 and 3). For (C), the RT-independent NCL events, which were supported by both AMV- and MMLV-based reads, were considered only (see the text). (D) Comparisons of percentages of tsRNA-involved events (i.e. TS-only and TS-circRNA events) that were detected in both poly(A)-selected data from hESCs and non-poly(A)-selected data from non-ESC samples. (E) Comparison of percentages of circRNA-involved events (i.e. circRNA-only and TS-circRNA events) that were detected in both non-poly(A)-selected data from hESCs and poly(A)-selected data from non-ESC samples. (F) qRT-PCR analyses of the expression fold changes for the selected 42 TS-circRNA events and two controls, GAPDH (which is poly(A)-tailed and must be degraded by RNase R treatment) and CDR1as (which is RNase R-resistant and must be non-polyadenylated) (1,5,6), before and after RNase R treatment in H9 hESCs. (G) Comparisons of expression fold changes for the selected 40 TS-circRNA events in poly(A)-tailed RNAs (oligo-dT pull down) and poly(A)-tailed RNAs treated with RNase R in H9 hESCs. The green (F) and blue (G) asterisks represent statistical significances of expression fold changes between the selected events and the controls (GAPDH for (F), green dashed lines; CDR1as for (G), blue dashed line). For (G), the red asterisks represent statistical significances of expression fold changes between the poly(A)-tailed RNAs and the poly(A)-tailed RNAs treated with RNase R. qRT-PCR experiments were performed in triplicate and repeated twice. Error bars represent the mean values ± one standard deviation. The statistical significance was evaluated using the two-tailed Fisher’s exact test (D and E) and the two-tailed t-test (F and G), respectively. *P < 0.05, **P < 0.01 and ***P < 0.001. (H) Detection of expanded length of tsRNAs by nanopore long reads. Blue and orange arrows indicate two different pairs of primers of the three examined TS-circRNA events, FARSA, HIPK3 and CAMSAP1. Empty rectangles represent the exons located outside the predicted circles of exonic circRNAs (the blue solid rectangles). E, exon.