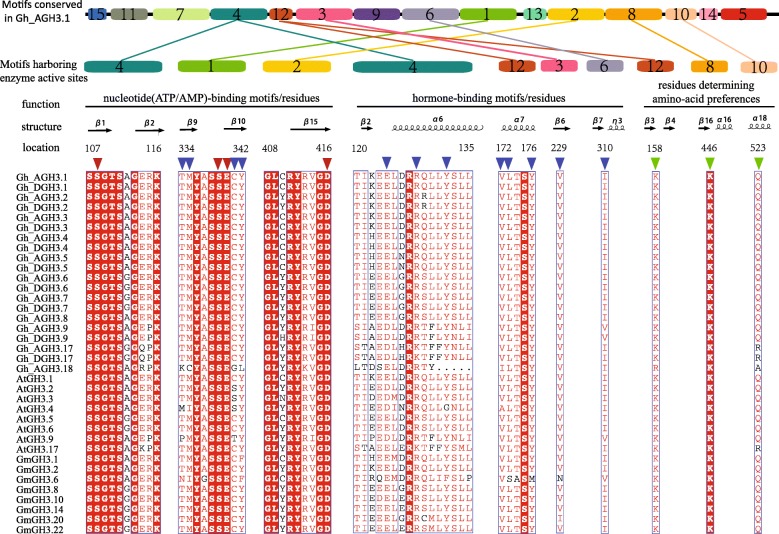
Fig. 3.
Sequence alignment of conserved residues or motifs. Protein sequences of 20 GhGH3s were aligned using MAFFT with all members of subfamily II in Arabidopsis and some in Soybean. The nucleotide (ATP/AMP) and hormone-binding motifs/residues as well as residues determining amino-acid preferences are conserved, indicating their similar functions as the IAA-amido synthetase in cotton. The colored triangles represent active sites and secondary structures near the active sites are also shown. Numbering at the top indicates the location of residues or motifs corresponding to Gh_AGH3.1. Motifs are put above the corresponding alignment of conserved residues and conserved motifs in full-length Gh_AGH3.1 are also shown as an example to display the locations of all motifs. The same motifs are linked by lines with the same color. The number in each motif indicates its code as described in Fig. 2c. Note that the ends of the lines do not indicate the actual positions of enzyme active sites